Visualization of retroviral envelope spikes in complex with the V3 loop antibody 447-52D on intact viruses by cryo-electron tomography
- PMID: 25122783
- PMCID: PMC4248906
- DOI: 10.1128/JVI.01596-14
Visualization of retroviral envelope spikes in complex with the V3 loop antibody 447-52D on intact viruses by cryo-electron tomography
Abstract
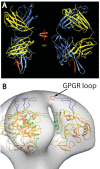
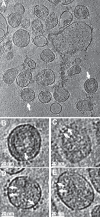
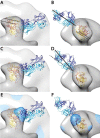
The gp120 portion of the envelope spike on human immunodeficiency virus type 1 (HIV-1) plays a critical role in viral entry into host cells and is a key target for the humoral immune response, and yet many structural details remain elusive. We have used cryoelectron tomography to visualize the binding of the broadly neutralizing monoclonal antibody (MAb) 447-52D to intact envelope spikes on virions of HIV-1 MN strain. Antibody 447-52D has previously been shown to bind to the tip of the V3 loop. Our results show antibody arms radiating from the sides of the gp120 protomers at a range of angles and place the antibody-bound V3 loop in an orientation that differs from that predicted by most current models but consistent with the idea that antibody binding dislodges the V3 loop from its location in the Env spike, making it flexible and disordered. These data reveal information on the position of the V3 loop and its relative flexibility and suggest that 447-52D neutralizes HIV-1 MN by capturing the V3 loop, blocking its interaction with the coreceptor and altering the structure of the envelope spike.
Importance: Antibody neutralization is one of the primary ways that the body fights infection with HIV. Because HIV is a highly mutable virus, the body must constantly produce new antibodies to counter new strains of HIV that the body itself is producing. Consequently, antibodies capable of neutralizing multiple HIV strains are comparatively few. An improved understanding of the mechanism of antibody neutralization might advance the development of immunogens. Most neutralizing antibodies target the Env glycoprotein spikes found on the virus surface. The broadly neutralizing antibody 447-52D targets the highly conserved β-turn of variable loop 3 (V3) of gp120. The importance of V3 lies in its contribution to the coreceptor binding site on the target cell. We show here that 447-52D binding to V3 converts the Env conformation from closed to open and makes the V3 loop highly flexible, implying disruption of coreceptor binding and attachment to the target cell.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures


Similar articles
-
Structure of Simian Immunodeficiency Virus Envelope Spikes Bound with CD4 and Monoclonal Antibody 36D5.J Virol. 2017 Jul 27;91(16):e00134-17. doi: 10.1128/JVI.00134-17. Print 2017 Aug 15. J Virol. 2017. PMID: 28539445 Free PMC article.
-
Design of immunogens that present the crown of the HIV-1 V3 loop in a conformation competent to generate 447-52D-like antibodies.Biochem J. 2006 Nov 1;399(3):483-91. doi: 10.1042/BJ20060588. Biochem J. 2006. PMID: 16827663 Free PMC article.
-
Functional and Structural Characterization of Human V3-Specific Monoclonal Antibody 2424 with Neutralizing Activity against HIV-1 JRFL.J Virol. 2015 Sep;89(17):9090-102. doi: 10.1128/JVI.01280-15. Epub 2015 Jun 24. J Virol. 2015. PMID: 26109728 Free PMC article.
-
Structural studies of human HIV-1 V3 antibodies.Hum Antibodies. 2005;14(3-4):73-80. Hum Antibodies. 2005. PMID: 16720977 Review.
-
Human antibodies that neutralize HIV-1: identification, structures, and B cell ontogenies.Immunity. 2012 Sep 21;37(3):412-25. doi: 10.1016/j.immuni.2012.08.012. Immunity. 2012. PMID: 22999947 Free PMC article. Review.
Cited by
-
Morphology and ultrastructure of retrovirus particles.AIMS Biophys. 2015;2(3):343-369. doi: 10.3934/biophy.2015.3.343. Epub 2015 Aug 18. AIMS Biophys. 2015. PMID: 26448965 Free PMC article.
-
Structure of Simian Immunodeficiency Virus Envelope Spikes Bound with CD4 and Monoclonal Antibody 36D5.J Virol. 2017 Jul 27;91(16):e00134-17. doi: 10.1128/JVI.00134-17. Print 2017 Aug 15. J Virol. 2017. PMID: 28539445 Free PMC article.
-
Design of HIV Coreceptor Derived Peptides That Inhibit Viral Entry at Submicromolar Concentrations.Mol Pharm. 2017 Aug 7;14(8):2681-2689. doi: 10.1021/acs.molpharmaceut.7b00155. Epub 2017 Jul 13. Mol Pharm. 2017. PMID: 28494151 Free PMC article.
-
Segmentation by classification: A novel and reliable approach for semi-automatic selection of HIV/SIV envelope spikes.J Struct Biol. 2020 Jan 1;209(1):107426. doi: 10.1016/j.jsb.2019.107426. Epub 2019 Nov 13. J Struct Biol. 2020. PMID: 31733279 Free PMC article.
-
Antibody Responses After Analytic Treatment Interruption in Human Immunodeficiency Virus-1-Infected Individuals on Early Initiated Antiretroviral Therapy.Open Forum Infect Dis. 2016 May 19;3(2):ofw100. doi: 10.1093/ofid/ofw100. eCollection 2016 Apr. Open Forum Infect Dis. 2016. PMID: 27419172 Free PMC article.
References
-
- Cohen J. 2003. Public health: AIDS vaccine trial produces disappointment and confusion. Science 299:1290–1291. - PubMed
-
- Lifson JD, Rossio JL, Piatak M, Jr, Bess J, Jr, Chertova E, Schneider DK, Coalter VJ, Poore B, Kiser RF, Imming RJ, Scarzello AJ, Henderson LE, Alvord WG, Hirsch VM, Benveniste RE, Arthur LO. 2004. Evaluation of the safety, immunogenicity, and protective efficacy of whole inactivated simian immunodeficiency virus (SIV) vaccines with conformationally and functionally intact envelope glycoproteins. AIDS Res. Hum. Retrovir. 20:772–787. 10.1089/0889222041524661. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

