7SL RNA represses p53 translation by competing with HuR
- PMID: 25123665
- PMCID: PMC4150789
- DOI: 10.1093/nar/gku686
7SL RNA represses p53 translation by competing with HuR
Abstract
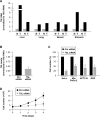
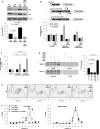
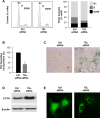
Noncoding RNAs (ncRNAs) and RNA-binding proteins are potent post-transcriptional regulators of gene expression. The ncRNA 7SL is upregulated in cancer cells, but its impact upon the phenotype of cancer cells is unknown. Here, we present evidence that 7SL forms a partial hybrid with the 3'-untranslated region (UTR) of TP53 mRNA, which encodes the tumor suppressor p53. The interaction of 7SL with TP53 mRNA reduced p53 translation, as determined by analyzing p53 expression levels, nascent p53 translation and TP53 mRNA association with polysomes. Silencing 7SL led to increased binding of HuR to TP53 mRNA, an interaction that led to the promotion of p53 translation and increased p53 abundance. We propose that the competition between 7SL and HuR for binding to TP53 3'UTR contributes to determining the magnitude of p53 translation, in turn affecting p53 levels and the growth-suppressive function of p53. Our findings suggest that targeting 7SL may be effective in the treatment of cancers with reduced p53 levels.
Published by Oxford University Press on behalf of Nucleic Acids Research 2014. This work is written by (a) US Government employee(s) and is in the public domain in the US.
Figures



References
-
- Faghihi M.A., Modarresi F., Khalil A.M., Wood D.E., Sahagan B.G., Morgan T.E., Finch C.E., St Laurent G., 3rd, Kenny P.J., Wahlestedt C. Expression of a noncoding RNA is elevated in Alzheimer's disease and drives rapid feed-forward regulation of beta-secretase. Nat. Med. 2008;14:723–730. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

