Bim controls IL-15 availability and limits engagement of multiple BH3-only proteins
- PMID: 25124553
- PMCID: PMC4262773
- DOI: 10.1038/cdd.2014.118
Bim controls IL-15 availability and limits engagement of multiple BH3-only proteins
Abstract
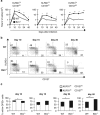
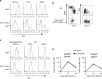
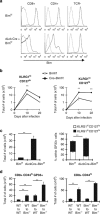
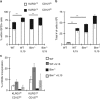
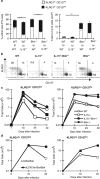
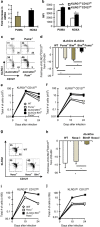
During the effector CD8+ T-cell response, transcriptional differentiation programs are engaged that promote effector T cells with varying memory potential. Although these differentiation programs have been used to explain which cells die as effectors and which cells survive and become memory cells, it is unclear if the lack of cell death enhances memory. Here, we investigated effector CD8+ T-cell fate in mice whose death program has been largely disabled because of the loss of Bim. Interestingly, the absence of Bim resulted in a significant enhancement of effector CD8+ T cells with more memory potential. Bim-driven control of memory T-cell development required T-cell-specific, but not dendritic cell-specific, expression of Bim. Both total and T-cell-specific loss of Bim promoted skewing toward memory precursors, by enhancing the survival of memory precursors, and limiting the availability of IL-15. Decreased IL-15 availability in Bim-deficient mice facilitated the elimination of cells with less memory potential via the additional pro-apoptotic molecules Noxa and Puma. Combined, these data show that Bim controls memory development by limiting the survival of pre-memory effector cells. Further, by preventing the consumption of IL-15, Bim limits the role of Noxa and Puma in causing the death of effector cells with less memory potential.
Figures

Similar articles
-
Bcl-2 allows effector and memory CD8+ T cells to tolerate higher expression of Bim.J Immunol. 2011 May 15;186(10):5729-37. doi: 10.4049/jimmunol.1100102. Epub 2011 Mar 30. J Immunol. 2011. PMID: 21451108 Free PMC article.
-
The BH3-only proteins Bim and Puma cooperate to impose deletional tolerance of organ-specific antigens.Immunity. 2012 Sep 21;37(3):451-62. doi: 10.1016/j.immuni.2012.05.030. Epub 2012 Sep 6. Immunity. 2012. PMID: 22960223 Free PMC article.
-
Puma cooperates with Bim, the rate-limiting BH3-only protein in cell death during lymphocyte development, in apoptosis induction.J Exp Med. 2006 Dec 25;203(13):2939-51. doi: 10.1084/jem.20061552. Epub 2006 Dec 18. J Exp Med. 2006. PMID: 17178918 Free PMC article.
-
Mechanisms of T cell death in the liver: to Bim or not to Bim?Dig Dis. 2010;28(1):14-24. doi: 10.1159/000282060. Epub 2010 May 7. Dig Dis. 2010. PMID: 20460886 Review.
-
Contracting the 'mus cells'--does down-sizing suit us for diving into the memory pool?Immunol Rev. 2010 Jul;236:54-67. doi: 10.1111/j.1600-065X.2010.00920.x. Immunol Rev. 2010. PMID: 20636808 Free PMC article. Review.
Cited by
-
Regulation of Bim in Health and Disease.Oncotarget. 2015 Sep 15;6(27):23058-134. doi: 10.18632/oncotarget.5492. Oncotarget. 2015. PMID: 26405162 Free PMC article. Review.
-
Dying to protect: cell death and the control of T-cell homeostasis.Immunol Rev. 2017 May;277(1):21-43. doi: 10.1111/imr.12538. Immunol Rev. 2017. PMID: 28462527 Free PMC article. Review.
-
Intracellular checkpoints for NK cell cancer immunotherapy.Front Med. 2024 Oct;18(5):763-777. doi: 10.1007/s11684-024-1090-6. Epub 2024 Sep 28. Front Med. 2024. PMID: 39340588 Review.
-
CCR7 expression alters memory CD8 T-cell homeostasis by regulating occupancy in IL-7- and IL-15-dependent niches.Proc Natl Acad Sci U S A. 2016 Jul 19;113(29):8278-83. doi: 10.1073/pnas.1602899113. Epub 2016 Jul 6. Proc Natl Acad Sci U S A. 2016. PMID: 27385825 Free PMC article.
-
Caspase-8-dependent control of NK- and T cell responses during cytomegalovirus infection.Med Microbiol Immunol. 2019 Aug;208(3-4):555-571. doi: 10.1007/s00430-019-00616-7. Epub 2019 May 16. Med Microbiol Immunol. 2019. PMID: 31098689 Review.
References
-
- Cannarile MA, Lind NA, Rivera R, Sheridan AD, Camfield KA, Wu BB, et al. Transcriptional regulator Id2 mediates CD8+ T cell immunity. Nat Immunol. 2006;7:1317–1325. - PubMed
-
- Kallies A, Xin A, Belz GT, Nutt SL. Blimp-1 transcription factor is required for the differentiation of effector CD8(+) T cells and memory responses. Immunity. 2009;31:283–295. - PubMed
-
- Intlekofer AM, Takemoto N, Wherry EJ, Longworth SA, Northrup JT, Palanivel VR, et al. Effector and memory CD8+ T cell fate coupled by T-bet and eomesodermin. Nat Immunol. 2005;6:1236–1244. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

