Genome-wide DNA methylation profiles in progression to in situ and invasive carcinoma of the breast with impact on gene transcription and prognosis
- PMID: 25146004
- PMCID: PMC4165906
- DOI: 10.1186/PREACCEPT-2333349012841587
Genome-wide DNA methylation profiles in progression to in situ and invasive carcinoma of the breast with impact on gene transcription and prognosis
Abstract
Background: Ductal carcinoma in situ (DCIS) of the breast is a precursor of invasive breast carcinoma. DNA methylation alterations are thought to be an early event in progression of cancer, and may prove valuable as a tool in clinical decision making and for understanding neoplastic development.
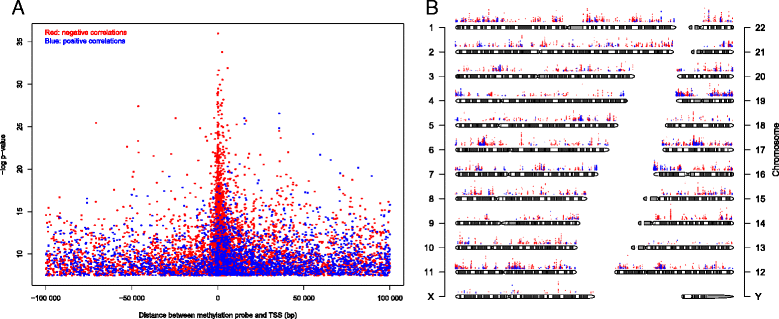
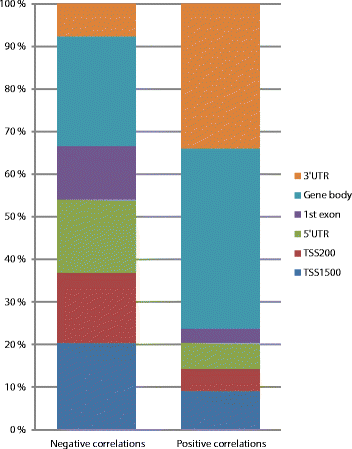
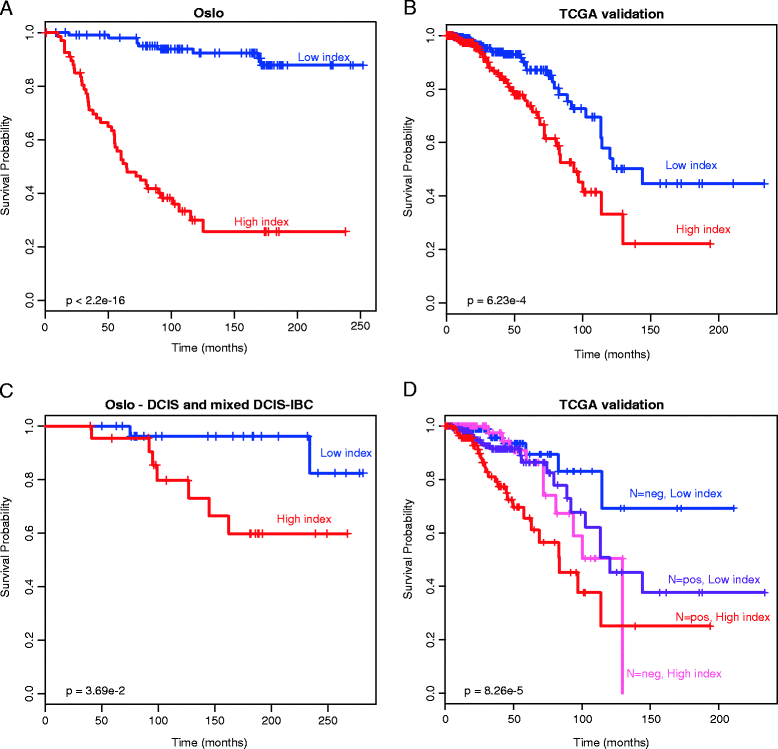
Results: We generate genome-wide DNA methylation profiles of 285 breast tissue samples representing progression of cancer, and validate methylation changes between normal and DCIS in an independent dataset of 15 normal and 40 DCIS samples. We also validate a prognostic signature on 583 breast cancer samples from The Cancer Genome Atlas. Our analysis reveals that DNA methylation profiles of DCIS are radically altered compared to normal breast tissue, involving more than 5,000 genes. Changes between DCIS and invasive breast carcinoma involve around 1,000 genes. In tumors, DNA methylation is associated with gene expression of almost 3,000 genes, including both negative and positive correlations. A prognostic signature based on methylation level of 18 CpGs is associated with survival of breast cancer patients with invasive tumors, as well as with survival of patients with DCIS and mixed lesions of DCIS and invasive breast carcinoma.
Conclusions: This work demonstrates that changes in the epigenome occur early in the neoplastic progression, provides evidence for the possible utilization of DNA methylation-based markers of progression in the clinic, and highlights the importance of epigenetic changes in carcinogenesis.
Figures

References
-
- Bediaga NG, Acha-Sagredo A, Guerra I, Viguri A, Albaina C, Ruiz DI, Rezola R, Alberdi MJ, Dopazo J, Montaner D, Renobales M, Fernandez AF, Field JK, Fraga MF, Liloglou T, de Pancorbo MM. DNA methylation epigenotypes in breast cancer molecular subtypes. Breast Cancer Res. 2010;12:R77. doi: 10.1186/bcr2721. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

