Early lineage restriction in temporally distinct populations of Mesp1 progenitors during mammalian heart development
- PMID: 25150979
- PMCID: PMC6984965
- DOI: 10.1038/ncb3024
Early lineage restriction in temporally distinct populations of Mesp1 progenitors during mammalian heart development
Abstract
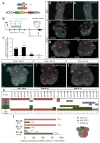
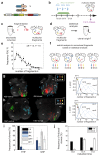
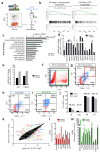
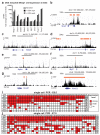
Cardiac development arises from two sources of mesoderm progenitors, the first heart field (FHF) and the second (SHF). Mesp1 has been proposed to mark the most primitive multipotent cardiac progenitors common for both heart fields. Here, using clonal analysis of the earliest prospective cardiovascular progenitors in a temporally controlled manner during early gastrulation, we found that Mesp1 progenitors consist of two temporally distinct pools of progenitors restricted to either the FHF or the SHF. FHF progenitors were unipotent, whereas SHF progenitors were either unipotent or bipotent. Microarray and single-cell PCR with reverse transcription analysis of Mesp1 progenitors revealed the existence of molecularly distinct populations of Mesp1 progenitors, consistent with their lineage and regional contribution. Together, these results provide evidence that heart development arises from distinct populations of unipotent and bipotent cardiac progenitors that independently express Mesp1 at different time points during their specification, revealing that the regional segregation and lineage restriction of cardiac progenitors occur very early during gastrulation.
Figures


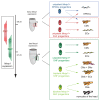
References
-
- Garry DJ, Olson EN. A common progenitor at the heart of development. Cell. 2006;127:1101–1104. - PubMed
-
- Kelly RG, Brown NA, Buckingham ME. The arterial pole of the mouse heart forms from Fgf10-expressing cells in pharyngeal mesoderm. Developmental cell. 2001;1:435–440. - PubMed
-
- Meilhac SM, Esner M, Kelly RG, Nicolas JF, Buckingham ME. The clonal origin of myocardial cells in different regions of the embryonic mouse heart. Developmental cell. 2004;6:685–698. - PubMed
-
- Saga Y, et al. MesP1 is expressed in the heart precursor cells and required for the formation of a single heart tube. Development (Cambridge, England) 1999;126:3437–3447. - PubMed
-
- Bondue A, Blanpain C. Mesp1: a key regulator of cardiovascular lineage commitment. Circ Res. 2010;107:1414–1427. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

