Fusion-related host proteins are actively regulated by NA during influenza infection as revealed by quantitative proteomics analysis
- PMID: 25153908
- PMCID: PMC4143309
- DOI: 10.1371/journal.pone.0105947
Fusion-related host proteins are actively regulated by NA during influenza infection as revealed by quantitative proteomics analysis
Abstract
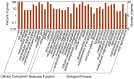
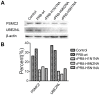
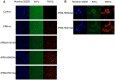
Three recombinant influenza A viruses with different neuraminidases (NAs) in the background of A/PR/8/34 (PR8), named rPR8-H5N1NA, rPR8-H9N2NA, and rPR8-H1N1NA, derived from H5N1, H9N2, H1N1 (swine) viruses, respectively, were constructed. We performed a quantitative proteomics analysis to investigate differential protein expression in Madin-Darby canine kidney (MDCK) cells infected with recombinant and wild-type influenza viruses to determine whether NA replacement would alter host cell gene expression. Using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF-TOF MS) and two-dimensional gel electrophoresis (2-DE), we identified 12 up-regulated and 49 down-regulated protein spots, including cytoskeletal proteins, molecular biosynthesis proteins, ubiquitin-proteasome pathway proteins, and heat shock proteins. The most significant changes in infected cells were observed for molecular biosynthesis proteins. We found more differentially expressed protein spots in cells infected with rPR8-H5N1NA or rPR8-H9N2NA viruses than cells infected with wild-type virus. Many of those proteins are postulated to be involved in cell-cell fusion, but the full mechanism remains to be explored. Meanwhile, our data demonstrate that the wild-type virus has evolutionary advantages over recombinant viruses.
Conflict of interest statement
Figures


Similar articles
-
NA proteins of influenza A viruses H1N1/2009, H5N1, and H9N2 show differential effects on infection initiation, virus release, and cell-cell fusion.PLoS One. 2013;8(1):e54334. doi: 10.1371/journal.pone.0054334. Epub 2013 Jan 22. PLoS One. 2013. PMID: 23349854 Free PMC article.
-
Effects of different polymerases of avian influenza viruses on the growth and pathogenicity of A/Puerto Rico/8/1934 (H1N1)-derived reassorted viruses.Vet Microbiol. 2014 Jan 10;168(1):41-9. doi: 10.1016/j.vetmic.2013.10.011. Epub 2013 Oct 26. Vet Microbiol. 2014. PMID: 24296300
-
Analysis of cellular proteome alterations in porcine alveolar macrophage cells infected with 2009 (H1N1) and classical swine H1N1 influenza viruses.J Proteomics. 2012 Mar 16;75(6):1732-41. doi: 10.1016/j.jprot.2011.12.012. Epub 2011 Dec 20. J Proteomics. 2012. PMID: 22202185
-
Influenza Neuraminidase Subtype N1: Immunobiological Properties and Functional Assays for Specific Antibody Response.PLoS One. 2016 Apr 7;11(4):e0153183. doi: 10.1371/journal.pone.0153183. eCollection 2016. PLoS One. 2016. PMID: 27054879 Free PMC article.
-
Neuraminidase inhibitor susceptibility and neuraminidase enzyme kinetics of human influenza A and B viruses circulating in Thailand in 2010-2015.PLoS One. 2018 Jan 11;13(1):e0190877. doi: 10.1371/journal.pone.0190877. eCollection 2018. PLoS One. 2018. PMID: 29324781 Free PMC article.
Cited by
-
Competitive Cooperation of Hemagglutinin and Neuraminidase during Influenza A Virus Entry.Viruses. 2019 May 20;11(5):458. doi: 10.3390/v11050458. Viruses. 2019. PMID: 31137516 Free PMC article. Review.
-
Proteomic analysis of differential expression of lung proteins in response to highly pathogenic avian influenza virus infection in chickens.Arch Virol. 2022 Jan;167(1):141-152. doi: 10.1007/s00705-021-05287-5. Epub 2021 Nov 16. Arch Virol. 2022. PMID: 34786609
-
High throughput discovery of protein variants using proteomics informed by transcriptomics.Nucleic Acids Res. 2018 Jun 1;46(10):4893-4902. doi: 10.1093/nar/gky295. Nucleic Acids Res. 2018. PMID: 29718325 Free PMC article.
-
Applied Proteomics in 'One Health'.Proteomes. 2021 Jun 30;9(3):31. doi: 10.3390/proteomes9030031. Proteomes. 2021. PMID: 34208880 Free PMC article. Review.
References
-
- Liu N, Song W, Wang P, Lee K, Chan W, et al. (2008) Proteomics analysis of differential expression of cellular proteins in response to avian H9N2 virus infection in human cells. Proteomics 8: 1851–1858. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

