Coupling mRNA synthesis and decay
- PMID: 25154419
- PMCID: PMC4248707
- DOI: 10.1128/MCB.00535-14
Coupling mRNA synthesis and decay
Abstract
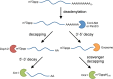
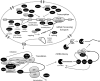
What has been will be again, what has been done will be done again; there is nothing new under the sun. -Ecclesiastes 1:9 (New International Version) Posttranscriptional regulation of gene expression has an important role in defining the phenotypic characteristics of an organism. Well-defined steps in mRNA metabolism that occur in the nucleus-capping, splicing, and polyadenylation-are mechanistically linked to the process of transcription. Recent evidence suggests another link between RNA polymerase II (Pol II) and a posttranscriptional process that occurs in the cytoplasm-mRNA decay. This conclusion appears to represent a conundrum. How could mRNA synthesis in the nucleus and mRNA decay in the cytoplasm be mechanistically linked? After a brief overview of mRNA processing, we will review the recent evidence for transcription-coupled mRNA decay and the possible involvement of Snf1, the Saccharomyces cerevisiae ortholog of AMP-activated protein kinase, in this process.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
