Near-real-time analysis of the phenotypic responses of Escherichia coli to 1-butanol exposure using Raman Spectroscopy
- PMID: 25157078
- PMCID: PMC4248880
- DOI: 10.1128/JB.01590-14
Near-real-time analysis of the phenotypic responses of Escherichia coli to 1-butanol exposure using Raman Spectroscopy
Abstract
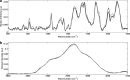
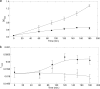
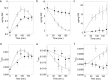
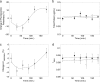
Raman spectroscopy was used to study the time course of phenotypic responses of Escherichia coli (DH5α) to 1-butanol exposure (1.2% [vol/vol]). Raman spectroscopy is of interest for bacterial phenotyping because it can be performed (i) in near real time, (ii) with minimal sample preparation (label-free), and (iii) with minimal spectral interference from water. Traditional off-line analytical methodologies were applied to both 1-butanol-treated and control cells to draw correlations with Raman data. Here, distinct sets of Raman bands are presented that characterize phenotypic traits of E. coli with maximized correlation to off-line measurements. In addition, the observed time course phenotypic responses of E. coli to 1.2% (vol/vol) 1-butanol exposure included the following: (i) decreased saturated fatty acids levels, (ii) retention of unsaturated fatty acids and low levels of cyclopropane fatty acids, (iii) increased membrane fluidity following the initial response of increased rigidity, and (iv) no changes in total protein content or protein-derived amino acid composition. For most phenotypic traits, correlation coefficients between Raman spectroscopy and traditional off-line analytical approaches exceeded 0.75, and major trends were captured. The results suggest that near-real-time Raman spectroscopy is suitable for approximating metabolic and physiological phenotyping of bacterial cells subjected to toxic environmental conditions.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures
Similar articles
-
Raman spectroscopy detects phenotypic differences among Escherichia coli enriched for 1-butanol tolerance using a metagenomic DNA library.Biotechnol J. 2016 Jul;11(7):877-89. doi: 10.1002/biot.201500144. Epub 2016 Mar 15. Biotechnol J. 2016. PMID: 26814030
-
Isolation of butanol- and isobutanol-tolerant bacteria and physiological characterization of their butanol tolerance.Appl Environ Microbiol. 2013 Nov;79(22):6998-7005. doi: 10.1128/AEM.02900-13. Epub 2013 Sep 6. Appl Environ Microbiol. 2013. PMID: 24014527 Free PMC article.
-
Phenotypic profiling of antibiotic response signatures in Escherichia coli using Raman spectroscopy.Antimicrob Agents Chemother. 2014;58(3):1302-14. doi: 10.1128/AAC.02098-13. Epub 2013 Dec 2. Antimicrob Agents Chemother. 2014. PMID: 24295982 Free PMC article.
-
Engineering Escherichia coli Cell Factories for n-Butanol Production.Adv Biochem Eng Biotechnol. 2016;155:141-63. doi: 10.1007/10_2015_306. Adv Biochem Eng Biotechnol. 2016. PMID: 25662903 Review.
-
Role of efflux in enhancing butanol tolerance of bacteria.J Biotechnol. 2020 Aug 20;320:17-27. doi: 10.1016/j.jbiotec.2020.06.008. Epub 2020 Jun 15. J Biotechnol. 2020. PMID: 32553531 Review.
Cited by
-
Discrimination of Stressed and Non-Stressed Food-Related Bacteria Using Raman-Microspectroscopy.Foods. 2022 May 22;11(10):1506. doi: 10.3390/foods11101506. Foods. 2022. PMID: 35627076 Free PMC article.
-
SERS-assisted characterization of cell biomass from biofilm-forming Acinetobacter baumannii strains using chemometric tools.RSC Adv. 2025 Feb 10;15(6):4581-4592. doi: 10.1039/d4ra06267a. eCollection 2025 Feb 6. RSC Adv. 2025. PMID: 39931412 Free PMC article.
-
Development and Automation of a Bacterial Biosensor to the Targeting of the Pollutants Toxic Effects by Portable Raman Spectrometer.Sensors (Basel). 2022 Jun 8;22(12):4352. doi: 10.3390/s22124352. Sensors (Basel). 2022. PMID: 35746134 Free PMC article.
-
Characterizing glucose, illumination, and nitrogen-deprivation phenotypes of Synechocystis PCC6803 with Raman spectroscopy.PeerJ. 2020 Mar 30;8:e8585. doi: 10.7717/peerj.8585. eCollection 2020. PeerJ. 2020. PMID: 32266111 Free PMC article.
-
Raman chemometric urinalysis (Rametrix) as a screen for bladder cancer.PLoS One. 2020 Aug 21;15(8):e0237070. doi: 10.1371/journal.pone.0237070. eCollection 2020. PLoS One. 2020. PMID: 32822394 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

