C-terminal domain swapping of SSB changes the size of the ssDNA binding site
- PMID: 25162017
- PMCID: PMC4137731
- DOI: 10.1155/2014/573936
C-terminal domain swapping of SSB changes the size of the ssDNA binding site
Abstract
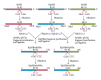
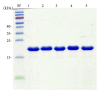
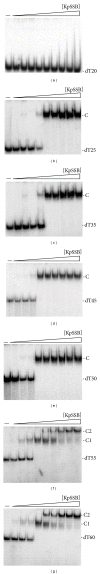
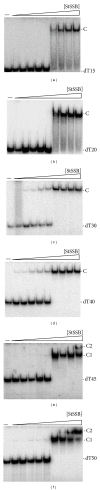
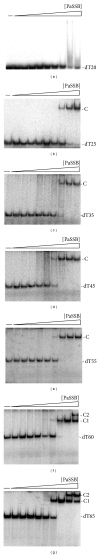
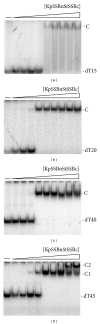
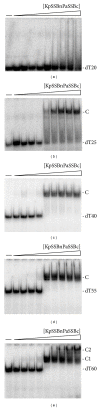
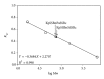
Single-stranded DNA-binding protein (SSB) plays an important role in DNA metabolism, including DNA replication, repair, and recombination, and is therefore essential for cell survival. Bacterial SSB consists of an N-terminal ssDNA-binding/oligomerization domain and a flexible C-terminal protein-protein interaction domain. We characterized the ssDNA-binding properties of Klebsiella pneumoniae SSB (KpSSB), Salmonella enterica Serovar Typhimurium LT2 SSB (StSSB), Pseudomonas aeruginosa PAO1 SSB (PaSSB), and two chimeric KpSSB proteins, namely, KpSSBnStSSBc and KpSSBnPaSSBc. The C-terminal domain of StSSB or PaSSB was exchanged with that of KpSSB through protein chimeragenesis. By using the electrophoretic mobility shift assay, we characterized the stoichiometry of KpSSB, StSSB, PaSSB, KpSSBnStSSBc, and KpSSBnPaSSBc, complexed with a series of ssDNA homopolymers. The binding site sizes were determined to be 26 ± 2, 21 ± 2, 29 ± 2, 21 ± 2, and 29 ± 2 nucleotides (nt), respectively. Comparison of the binding site sizes of KpSSB, KpSSBnStSSBc, and KpSSBnPaSSBc showed that the C-terminal domain swapping of SSB changes the size of the binding site. Our observations suggest that not only the conserved N-terminal domain but also the C-terminal domain of SSB is an important determinant for ssDNA binding.
Figures

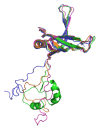
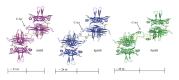
Similar articles
-
Characterization of a single-stranded DNA binding protein from Salmonella enterica serovar Typhimurium LT2.Protein J. 2011 Feb;30(2):102-8. doi: 10.1007/s10930-011-9309-1. Protein J. 2011. PMID: 21267641
-
Characterization of a single-stranded DNA-binding protein from Pseudomonas aeruginosa PAO1.Protein J. 2011 Jan;30(1):20-6. doi: 10.1007/s10930-010-9297-6. Protein J. 2011. PMID: 21132356
-
Characterization of a single-stranded DNA-binding protein from Klebsiella pneumoniae: mutation at either Arg73 or Ser76 causes a less cooperative complex on DNA.Genes Cells. 2012 Feb;17(2):146-57. doi: 10.1111/j.1365-2443.2011.01577.x. Genes Cells. 2012. PMID: 22244199
-
The mechanism of action of the SSB interactome reveals it is the first OB-fold family of genome guardians in prokaryotes.Protein Sci. 2021 Sep;30(9):1757-1775. doi: 10.1002/pro.4140. Epub 2021 Jun 14. Protein Sci. 2021. PMID: 34089559 Free PMC article. Review.
-
Single-Stranded DNA-Binding Proteins in the Archaea.Methods Mol Biol. 2021;2281:23-47. doi: 10.1007/978-1-0716-1290-3_2. Methods Mol Biol. 2021. PMID: 33847950 Review.
Cited by
-
Impact of ROS-Induced Damage of TCA Cycle Enzymes on Metabolism and Virulence of Salmonella enterica serovar Typhimurium.Front Microbiol. 2019 Apr 24;10:762. doi: 10.3389/fmicb.2019.00762. eCollection 2019. Front Microbiol. 2019. PMID: 31105651 Free PMC article.
-
SAAV2152 is a single-stranded DNA binding protein: the third SSB in Staphylococcus aureus.Oncotarget. 2018 Feb 5;9(29):20239-20254. doi: 10.18632/oncotarget.24427. eCollection 2018 Apr 17. Oncotarget. 2018. PMID: 29755648 Free PMC article.
-
Staphylococcus aureus single-stranded DNA-binding protein SsbA can bind but cannot stimulate PriA helicase.PLoS One. 2017 Jul 27;12(7):e0182060. doi: 10.1371/journal.pone.0182060. eCollection 2017. PLoS One. 2017. PMID: 28750050 Free PMC article.
-
The Inhibitory Effects and Cytotoxic Activities of the Stem Extract of Sarracenia purpurea against Melanoma Cells and the SsbA Protein.Plants (Basel). 2022 Nov 18;11(22):3164. doi: 10.3390/plants11223164. Plants (Basel). 2022. PMID: 36432892 Free PMC article.
-
Inhibition of a Putative Dihydropyrimidinase from Pseudomonas aeruginosa PAO1 by Flavonoids and Substrates of Cyclic Amidohydrolases.PLoS One. 2015 May 19;10(5):e0127634. doi: 10.1371/journal.pone.0127634. eCollection 2015. PLoS One. 2015. PMID: 25993634 Free PMC article.
References
-
- Richard DJ, Bolderson E, Khanna KK. Multiple human single-stranded DNA binding proteins function in genome maintenance: structural, biochemical and functional analysis. Critical Reviews in Biochemistry and Molecular Biology. 2009;44(2-3):98–116. - PubMed
-
- Richard DJ, Bolderson E, Cubeddu L, et al. Single-stranded DNA-binding protein hSSB1 is critical for genomic stability. Nature. 2008;453(7195):677–681. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

