Cytoscape: the network visualization tool for GenomeSpace workflows
- PMID: 25165537
- PMCID: PMC4133763
- DOI: 10.12688/f1000research.4492.2
Cytoscape: the network visualization tool for GenomeSpace workflows
Abstract
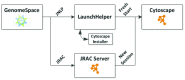
Modern genomic analysis often requires workflows incorporating multiple best-of-breed tools. GenomeSpace is a web-based visual workbench that combines a selection of these tools with mechanisms that create data flows between them. One such tool is Cytoscape 3, a popular application that enables analysis and visualization of graph-oriented genomic networks. As Cytoscape runs on the desktop, and not in a web browser, integrating it into GenomeSpace required special care in creating a seamless user experience and enabling appropriate data flows. In this paper, we present the design and operation of the Cytoscape GenomeSpace app, which accomplishes this integration, thereby providing critical analysis and visualization functionality for GenomeSpace users. It has been downloaded over 850 times since the release of its first version in September, 2013.
Conflict of interest statement
Figures
References
-
- Altintas I, Wang J, Crawl D, et al. : Challenges and approaches for distributed workflowdriven analysis of large-scale biological data. Proceedings of the Workshop on Data analytics in the Cloud at EDBT/ICDT 2012 Conference, DanaC2012,2012;73–78 10.1145/2320765.2320791 - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials