Gene transfers shaped the evolution of de novo NAD+ biosynthesis in eukaryotes
- PMID: 25169983
- PMCID: PMC4217691
- DOI: 10.1093/gbe/evu185
Gene transfers shaped the evolution of de novo NAD+ biosynthesis in eukaryotes
Abstract
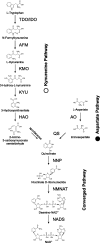
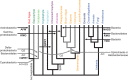
NAD(+) is an essential molecule for life, present in each living cell. It can function as an electron carrier or cofactor in redox biochemistry and energetics, and serves as substrate to generate the secondary messenger cyclic ADP ribose and nicotinic acid adenine dinucleotide phosphate. Although de novo NAD(+) biosynthesis is essential, different metabolic pathways exist in different eukaryotic clades. The kynurenine pathway starting with tryptophan was most likely present in the last common ancestor of all eukaryotes, and is active in fungi and animals. The aspartate pathway, detected in most photosynthetic eukaryotes, was probably acquired from the cyanobacterial endosymbiont that gave rise to chloroplasts. An evolutionary analysis of enzymes catalyzing de novo NAD(+) biosynthesis resulted in evolutionary trees incongruent with established organismal phylogeny, indicating numerous gene transfers. Endosymbiotic gene transfers probably introduced the aspartate pathway into eukaryotes and may have distributed it among different photosynthetic clades. In addition, several horizontal gene transfers substituted eukaryotic genes with bacterial orthologs. Although horizontal gene transfer is accepted as a key mechanism in prokaryotic evolution, it is supposed to be rare in eukaryotic evolution. The essential metabolic pathway of de novo NAD(+) biosynthesis in eukaryotes was shaped by numerous gene transfers.
Keywords: NAD+ biosynthesis; endosymbiotic gene transfer; horizontal gene transfer; metabolism.
© The Author(s) 2014. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures





References
-
- Abascal F, Zardoya R, Posada D. ProtTest: selection of best-fit models of protein evolution. Bioinformatics. 2005;21:2104–2105. - PubMed
-
- Altekar G, Dwarkadas S, Huelsenbeck JP, Ronquist F. Parallel metropolis coupled Markov chain Monte Carlo for Bayesian phylogenetic inference. Bioinformatics. 2004;20:407–415. - PubMed
-
- Andersson JO. Evolution of patchily distributed proteins shared between eukaryotes and prokaryotes: Dictyostelium as a case study. J Mol Microbiol Biotechnol. 2011;20:83–95. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

