Evidence for the retention of two evolutionary distinct plastids in dinoflagellates with diatom endosymbionts
- PMID: 25172904
- PMCID: PMC4217693
- DOI: 10.1093/gbe/evu182
Evidence for the retention of two evolutionary distinct plastids in dinoflagellates with diatom endosymbionts
Abstract
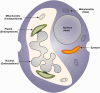
Dinoflagellates harboring diatom endosymbionts (termed "dinotoms") have undergone a process often referred to as "tertiary endosymbiosis"--the uptake of algae containing secondary plastids and integration of those plastids into the new host. In contrast to other tertiary plastids, and most secondary plastids, the endosymbiont of dinotoms is distinctly less reduced, retaining a number of cellular features, such as their nucleus and mitochondria and others, in addition to their plastid. This has resulted in redundancy between host and endosymbiont, at least between some mitochondrial and cytosolic metabolism, where this has been investigated. The question of plastidial redundancy is particularly interesting as the fate of the host dinoflagellate plastid is unclear. The host cytosol possesses an eyespot that has been postulated to be a remnant of the ancestral peridinin plastid, but this has not been tested, nor has its possible retention of plastid functions. To investigate this possibility, we searched for plastid-associated pathways and functions in transcriptomic data sets from three dinotom species. We show that the dinoflagellate host has indeed retained genes for plastid-associated pathways and that these genes encode targeting peptides similar to those of other dinoflagellate plastid-targeted proteins. Moreover, we also identified one gene encoding an essential component of the dinoflagellate plastid protein import machinery, altogether suggesting the presence of a functioning plastid import system in the host, and by extension a relict plastid. The presence of the same plastid-associated pathways in the endosymbiont also extends the known functional redundancy in dinotoms, further confirming the unusual state of plastid integration in this group of dinoflagellates.
Keywords: dinotom; redundancy; relict plastid; tertiary endosymbiosis.
© The Author(s) 2014. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
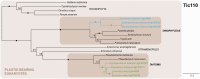
Figures


Similar articles
-
Functional Relationship between a Dinoflagellate Host and Its Diatom Endosymbiont.Mol Biol Evol. 2016 Sep;33(9):2376-90. doi: 10.1093/molbev/msw109. Epub 2016 Jun 13. Mol Biol Evol. 2016. PMID: 27297471
-
Dinotoms possess two evolutionary distinct autophagy-related ubiquitin-like conjugation systems.Protist. 2024 Dec;175(6):126067. doi: 10.1016/j.protis.2024.126067. Epub 2024 Sep 21. Protist. 2024. PMID: 39341116
-
A phylogenetic mosaic plastid proteome and unusual plastid-targeting signals in the green-colored dinoflagellate Lepidodinium chlorophorum.BMC Evol Biol. 2010 Jun 21;10:191. doi: 10.1186/1471-2148-10-191. BMC Evol Biol. 2010. PMID: 20565933 Free PMC article.
-
Integration of plastids with their hosts: Lessons learned from dinoflagellates.Proc Natl Acad Sci U S A. 2015 Aug 18;112(33):10247-54. doi: 10.1073/pnas.1421380112. Epub 2015 May 20. Proc Natl Acad Sci U S A. 2015. PMID: 25995366 Free PMC article. Review.
-
The endosymbiotic origin, diversification and fate of plastids.Philos Trans R Soc Lond B Biol Sci. 2010 Mar 12;365(1541):729-48. doi: 10.1098/rstb.2009.0103. Philos Trans R Soc Lond B Biol Sci. 2010. PMID: 20124341 Free PMC article. Review.
Cited by
-
Function and evolutionary origin of unicellular camera-type eye structure.PLoS One. 2015 Mar 3;10(3):e0118415. doi: 10.1371/journal.pone.0118415. eCollection 2015. PLoS One. 2015. PMID: 25734540 Free PMC article.
-
Nitzschia anatoliensis sp. nov., a cryptic diatom species from the highly alkaline Van Lake (Turkey).PeerJ. 2021 Oct 22;9:e12220. doi: 10.7717/peerj.12220. eCollection 2021. PeerJ. 2021. PMID: 34733585 Free PMC article.
-
Single-cell transcriptomics using spliced leader PCR: Evidence for multiple losses of photosynthesis in polykrikoid dinoflagellates.BMC Genomics. 2015 Jul 17;16(1):528. doi: 10.1186/s12864-015-1636-8. BMC Genomics. 2015. PMID: 26183220 Free PMC article.
-
Dinoflagellates with relic endosymbiont nuclei as models for elucidating organellogenesis.Proc Natl Acad Sci U S A. 2020 Mar 10;117(10):5364-5375. doi: 10.1073/pnas.1911884117. Epub 2020 Feb 24. Proc Natl Acad Sci U S A. 2020. PMID: 32094181 Free PMC article.
-
In the beginning was the word: How terminology drives our understanding of endosymbiotic organelles.Microb Cell. 2019 Jan 21;6(2):134-141. doi: 10.15698/mic2019.02.669. Microb Cell. 2019. PMID: 30740458 Free PMC article.
References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Apt KE, et al. In vivo characterization of diatom multipartite plastid targeting signals. J Cell Sci. 2002;115:4061–4069. - PubMed
-
- Bendtsen JD, Nielsen H, von Heijne G, Brunak S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol. 2004;340:783–795. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

