Thiol peroxidase deficiency leads to increased mutational load and decreased fitness in Saccharomyces cerevisiae
- PMID: 25173844
- PMCID: PMC4224179
- DOI: 10.1534/genetics.114.169243
Thiol peroxidase deficiency leads to increased mutational load and decreased fitness in Saccharomyces cerevisiae
Abstract
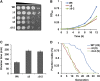
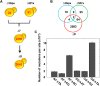
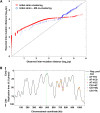
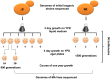
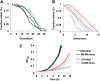
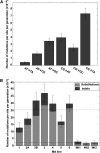
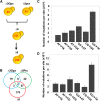
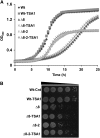
Thiol peroxidases are critical enzymes in the redox control of cellular processes that function by reducing low levels of hydroperoxides and regulating redox signaling. These proteins were also shown to regulate genome stability, but how their dysfunction affects the actual mutations in the genome is not known. Saccharomyces cerevisiae has eight thiol peroxidases of glutathione peroxidase and peroxiredoxin families, and the mutant lacking all these genes (∆8) is viable. In this study, we employed two independent ∆8 isolates to analyze the genome-wide mutation spectrum that results from deficiency in these enzymes. Deletion of these genes was accompanied by a dramatic increase in point mutations, many of which clustered in close proximity and scattered throughout the genome, suggesting strong mutational bias. We further subjected multiple lines of wild-type and ∆8 cells to long-term mutation accumulation, followed by genome sequencing and phenotypic characterization. ∆8 lines showed a significant increase in nonrecurrent point mutations and indels. The original ∆8 cells exhibited reduced growth rate and decreased life span, which were further reduced in all ∆8 mutation accumulation lines. Although the mutation spectrum of the two independent isolates was different, similar patterns of gene expression were observed, suggesting the direct contribution of thiol peroxidases to the observed phenotypes. Expression of a single thiol peroxidase could partially restore the growth phenotype of ∆8 cells. This study shows how deficiency in nonessential, yet critical and conserved oxidoreductase function, leads to increased mutational load and decreased fitness.
Keywords: Saccharomyces cerevisiae; genome stability; mutation; thiol peroxidase.
Copyright © 2014 by the Genetics Society of America.
Figures

Similar articles
-
Evidence that mutation accumulation does not cause aging in Saccharomyces cerevisiae.Aging Cell. 2015 Jun;14(3):366-71. doi: 10.1111/acel.12290. Epub 2015 Feb 22. Aging Cell. 2015. PMID: 25702753 Free PMC article.
-
Adaptive aneuploidy protects against thiol peroxidase deficiency by increasing respiration via key mitochondrial proteins.Proc Natl Acad Sci U S A. 2015 Aug 25;112(34):10685-90. doi: 10.1073/pnas.1505315112. Epub 2015 Aug 10. Proc Natl Acad Sci U S A. 2015. PMID: 26261310 Free PMC article.
-
Thiol peroxidases mediate specific genome-wide regulation of gene expression in response to hydrogen peroxide.Proc Natl Acad Sci U S A. 2011 Feb 15;108(7):2729-34. doi: 10.1073/pnas.1010721108. Epub 2011 Jan 31. Proc Natl Acad Sci U S A. 2011. PMID: 21282621 Free PMC article.
-
Thermosensitive phenotype of yeast mutant lacking thioredoxin peroxidase.Arch Biochem Biophys. 1998 Nov 1;359(1):99-106. doi: 10.1006/abbi.1998.0896. Arch Biochem Biophys. 1998. PMID: 9799566
-
Nuclear thioredoxin peroxidase Dot5 in Saccharomyces cerevisiae: roles in oxidative stress response and disruption of telomeric silencing.Appl Microbiol Biotechnol. 2004 Mar;64(1):120-4. doi: 10.1007/s00253-003-1421-5. Epub 2003 Aug 19. Appl Microbiol Biotechnol. 2004. PMID: 12925864
Cited by
-
ROS-Activated Ion Channels in Plants: Biophysical Characteristics, Physiological Functions and Molecular Nature.Int J Mol Sci. 2018 Apr 23;19(4):1263. doi: 10.3390/ijms19041263. Int J Mol Sci. 2018. PMID: 29690632 Free PMC article. Review.
-
Which Is the Most Significant Cause of Aging?Antioxidants (Basel). 2015 Dec 17;4(4):793-810. doi: 10.3390/antiox4040793. Antioxidants (Basel). 2015. PMID: 26783959 Free PMC article. Review.
-
Evidence that mutation accumulation does not cause aging in Saccharomyces cerevisiae.Aging Cell. 2015 Jun;14(3):366-71. doi: 10.1111/acel.12290. Epub 2015 Feb 22. Aging Cell. 2015. PMID: 25702753 Free PMC article.
-
Molecular signatures of aneuploidy-driven adaptive evolution.Nat Commun. 2020 Jan 30;11(1):588. doi: 10.1038/s41467-019-13669-2. Nat Commun. 2020. PMID: 32001709 Free PMC article.
-
Adaptive aneuploidy protects against thiol peroxidase deficiency by increasing respiration via key mitochondrial proteins.Proc Natl Acad Sci U S A. 2015 Aug 25;112(34):10685-90. doi: 10.1073/pnas.1505315112. Epub 2015 Aug 10. Proc Natl Acad Sci U S A. 2015. PMID: 26261310 Free PMC article.
References
-
- Avery A. M., Avery S. V., 2001. Saccharomyces cerevisiae expresses three phospholipid hydroperoxide glutathione peroxidases. J. Biol. Chem. 276: 33730–33735. - PubMed
-
- Brigelius-Flohe R., 1999. Tissue-specific functions of individual glutathione peroxidases. Free Radic. Biol. Med. 27: 951–965. - PubMed
-
- Brigelius-Flohe R., 2006. Glutathione peroxidases and redox-regulated transcription factors. Biol. Chem. 387: 1329–1335. - PubMed
-
- Cooke M. S., Evans M. D., Dizdaroglu M., Lunec J., 2003. Oxidative DNA damage: mechanisms, mutation, and disease. FASEB J. 17: 1195–1214. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

