Metabolomics in the fight against malaria
- PMID: 25185001
- PMCID: PMC4156452
- DOI: 10.1590/0074-0276140043
Metabolomics in the fight against malaria
Abstract
Metabolomics uses high-resolution mass spectrometry to provide a chemical fingerprint of thousands of metabolites present in cells, tissues or body fluids. Such metabolic phenotyping has been successfully used to study various biologic processes and disease states. High-resolution metabolomics can shed new light on the intricacies of host-parasite interactions in each stage of the Plasmodium life cycle and the downstream ramifications on the host's metabolism, pathogenesis and disease. Such data can become integrated with other large datasets generated using top-down systems biology approaches and be utilised by computational biologists to develop and enhance models of malaria pathogenesis relevant for identifying new drug targets or intervention strategies. Here, we focus on the promise of metabolomics to complement systems biology approaches in the quest for novel interventions in the fight against malaria. We introduce the Malaria Host-Pathogen Interaction Center (MaHPIC), a new systems biology research coalition. A primary goal of the MaHPIC is to generate systems biology datasets relating to human and non-human primate (NHP) malaria parasites and their hosts making these openly available from an online relational database. Metabolomic data from NHP infections and clinical malaria infections from around the world will comprise a unique global resource.
Figures
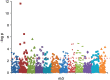
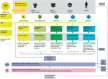
References
-
- Ariey F, Witkowski B, Amaratunga C, Beghain J, Langlois A-C, Khim N, Kim S, Duru V, Bouchier C, Ma L, Lim P, Leang R, Duong S, Sreng S, Suon S, Chuor CM, Bout DM, Ménard S, Rogers WO, Genton B, Fandeur T, Miotto O, Ringwald P, Le Bras J, Berry A, Barale J-C, Fairhurst RM, Benoit-Vical F, Mercereau-Puijalon O, Ménard D. A molecular marker of artemisinin-resistant Plasmodium falciparum malaria. Nature. 2014;505:50–55. - PMC - PubMed
-
- Aurrecoechea C, Brestelli J, Brunk BP, Dommer J, Fischer S, Gajria B, Gao X, Gingle A, Grant G, Harb OS, Heiges M, Innamorato F, Iodice J, Kissinger JC, Kraemer E, Li W, Miller JA, Nayak V, Pennington C, Pinney DF, Roos DS, Ross C, Stoeckert CJ, Treatman C, Wang H. PlasmoDB: a functional genomic database for malaria parasites. Nucleic Acids Res. 2009;37:D539–D543. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

