Progress in lactic acid bacterial phage research
- PMID: 25185514
- PMCID: PMC4155818
- DOI: 10.1186/1475-2859-13-S1-S1
Progress in lactic acid bacterial phage research
Abstract
Research on lactic acid bacteria (LAB) has advanced significantly over the past number of decades and these developments have been driven by the parallel advances in technologies such as genomics, bioinformatics, protein expression systems and structural biology, combined with the ever increasing commercial relevance of this group of microorganisms. Some of the more significant and impressive outputs have been in the domain of bacteriophage-host interactions which provides a prime example of the cutting-edge model systems represented by LAB research. Here, we present a retrospective overview of the key advances in LAB phage research including phage-host interactions and co-evolution. We describe how in many instances this knowledge can be pivotal in creating real improvements in the application of LAB cultures in commercial practice.
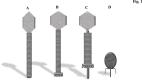
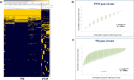
Figures



Similar articles
-
Physiological Role of Two-Component Signal Transduction Systems in Food-Associated Lactic Acid Bacteria.Adv Appl Microbiol. 2017;99:1-51. doi: 10.1016/bs.aambs.2016.12.002. Epub 2017 Jan 18. Adv Appl Microbiol. 2017. PMID: 28438266 Review.
-
Bacteriophage defence systems in lactic acid bacteria.Antonie Van Leeuwenhoek. 1999 Jul-Nov;76(1-4):89-113. Antonie Van Leeuwenhoek. 1999. PMID: 10532374 Review.
-
Towards metagenome-scale models for industrial applications--the case of Lactic Acid Bacteria.Curr Opin Biotechnol. 2013 Apr;24(2):200-6. doi: 10.1016/j.copbio.2012.11.003. Epub 2012 Nov 28. Curr Opin Biotechnol. 2013. PMID: 23200025 Review.
-
Lactic acid bacteria: life after genomics.Microb Biotechnol. 2011 May;4(3):318-22. doi: 10.1111/j.1751-7915.2011.00262.x. Microb Biotechnol. 2011. PMID: 21518298 Free PMC article. No abstract available.
-
Bacteriocins of lactic acid bacteria: extending the family.Appl Microbiol Biotechnol. 2016 Apr;100(7):2939-51. doi: 10.1007/s00253-016-7343-9. Epub 2016 Feb 10. Appl Microbiol Biotechnol. 2016. PMID: 26860942 Free PMC article. Review.
Cited by
-
Comparative Genomic Analysis Reveals Intestinal Habitat Adaptation of Ligilactobacillus equi Rich in Prophage and Degrading Cellulase.Molecules. 2022 Mar 14;27(6):1867. doi: 10.3390/molecules27061867. Molecules. 2022. PMID: 35335231 Free PMC article.
-
Host-encoded, cell surface-associated exopolysaccharide required for adsorption and infection by lactococcal P335 phage subtypes.Front Microbiol. 2022 Oct 4;13:971166. doi: 10.3389/fmicb.2022.971166. eCollection 2022. Front Microbiol. 2022. PMID: 36267184 Free PMC article.
-
Metabiotics Signature through Genome Sequencing and In Vitro Inhibitory Assessment of a Novel Lactococcus lactis Strain UTNCys6-1 Isolated from Amazonian Camu-Camu Fruits.Int J Mol Sci. 2023 Mar 24;24(7):6127. doi: 10.3390/ijms24076127. Int J Mol Sci. 2023. PMID: 37047101 Free PMC article.
-
Lactococcus lactis type III-A CRISPR-Cas system cleaves bacteriophage RNA.RNA Biol. 2019 Apr;16(4):461-468. doi: 10.1080/15476286.2018.1502589. Epub 2018 Oct 2. RNA Biol. 2019. PMID: 30081743 Free PMC article.
-
A Specific Sugar Moiety in the Lactococcus lactis Cell Wall Pellicle Is Required for Infection by CHPC971, a Member of the Rare 1706 Phage Species.Appl Environ Microbiol. 2019 Sep 17;85(19):e01224-19. doi: 10.1128/AEM.01224-19. Print 2019 Oct 1. Appl Environ Microbiol. 2019. PMID: 31350317 Free PMC article.
References
-
- Klaenhammer TR, Sanozky RB. Conjugal transfer from Streptococcus lactis ME2 of plasmids encoding phage resistance, nisin resistance and lactose-fermenting ability: evidence for a high-frequency conjugative plasmid responsible for abortive infection of virulent bacteriophage. J Gen Microbiol. 1985;131:1531–1541. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

