Salinity and bacterial diversity: to what extent does the concentration of salt affect the bacterial community in a saline soil?
- PMID: 25188357
- PMCID: PMC4154724
- DOI: 10.1371/journal.pone.0106662
Salinity and bacterial diversity: to what extent does the concentration of salt affect the bacterial community in a saline soil?
Erratum in
- PLoS One. 2014;9(11):e114658
Abstract
In this study, the evaluation of soil characteristics was coupled with a pyrosequencing analysis of the V2-V3 16S rRNA gene region in order to investigate the bacterial community structure and diversity in the A horizon of a natural saline soil located in Sicily (Italy). The main aim of the research was to assess the organisation and diversity of microbial taxa using a spatial scale that revealed physical and chemical heterogeneity of the habitat under investigation. The results provided information on the type of distribution of different bacterial groups as a function of spatial gradients of soil salinity and pH. The analysis of bacterial 16S rRNA showed differences in bacterial composition and diversity due to a variable salt concentration in the soil. The bacterial community showed a statistically significant spatial variability. Some bacterial phyla appeared spread in the whole area, whatever the salinity gradient. It emerged therefore that a patchy saline soil can not contain just a single microbial community selected to withstand extreme osmotic phenomena, but many communities that can be variously correlated to one or more environmental parameters. Sequences have been deposited to the SRA database and can be accessed on ID Project PRJNA241061.
Conflict of interest statement
Figures
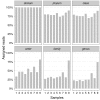

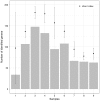

 Where Xij is the occurrence of the phyla ‘j’ in the site ‘i’ and N is the number of site in the dataset (in this case 9). Using this transformation each phyla assignment can be compared in all sites independently from its order of magnitude.
Where Xij is the occurrence of the phyla ‘j’ in the site ‘i’ and N is the number of site in the dataset (in this case 9). Using this transformation each phyla assignment can be compared in all sites independently from its order of magnitude.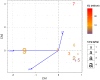



References
-
- Richards LA (1954) Diagnosis and improvement of saline and alkali soils. US Salinity Lab., US Department of Agriculture Handbook 60 California, USA.
-
- IUSS Working Group WRB (2007) World Reference Base for Soil Resources 2006, first update. 2007. World Soil Resources Reports No. 103. FAO, Rome.
-
- Soil Survey Staff (2010) Keys to Soil Taxonomy, 11th ed. USDA-Natural Resources Conservation Service, Washington, DC.
-
- Mavi MS, Sandarman J, Chittleborough DJ, Cox JW, Marchner P (2012) Sorption of dissolved organic matter in salt-affected soils: Effect of salinity, sodicity and texture. Sci. Total Environ. 435–436. - PubMed
-
- Rietz DN, Haynes RJ (2003) Effects of irrigation induced salinity and sodicity on soil microbial activity. Soil Biol. Biochem 35: 845–854.
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources

