Bällchen participates in proliferation control and prevents the differentiation of Drosophila melanogaster neuronal stem cells
- PMID: 25190057
- PMCID: PMC4197436
- DOI: 10.1242/bio.20148631
Bällchen participates in proliferation control and prevents the differentiation of Drosophila melanogaster neuronal stem cells
Abstract
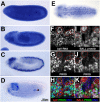
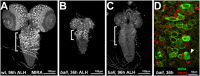
Stem cells continuously generate differentiating daughter cells and are essential for tissue homeostasis and development. Their capacity to self-renew as undifferentiated and actively dividing cells is controlled by either external signals from a cellular environment, the stem cell niche, or asymmetric distribution of cell fate determinants during cell division. Here we report that the protein kinase Bällchen (BALL) is required to prevent differentiation as well as to maintain normal proliferation of neuronal stem cells of Drosophila melanogaster, called neuroblasts. Our results show that the brains of ball mutant larvae are severely reduced in size, which is caused by a reduced proliferation rate of the neuroblasts. Moreover, ball mutant neuroblasts gradually lose the expression of the neuroblast determinants Miranda and aPKC, suggesting their premature differentiation. Our results indicate that BALL represents a novel cell intrinsic factor with a dual function regulating the proliferative capacity and the differentiation status of neuronal stem cells during development.
Keywords: Bällchen; Drosophila; Neuroblasts; Stem cells.
© 2014. Published by The Company of Biologists Ltd.
Conflict of interest statement
Figures


References
-
- Berger C., Harzer H., Burkard T. R., Steinmann J., van der Horst S., Laurenson A.-S., Novatchkova M., Reichert H., Knoblich J. A. (2012). FACS purification and transcriptome analysis of Drosophila neural stem cells reveals a role for Klumpfuss in self-renewal. Cell Reports 2, 407–418 10.1016/j.celrep.2012.07.008 - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

