Natural selection on coding and noncoding DNA sequences is associated with virulence genes in a plant pathogenic fungus
- PMID: 25193312
- PMCID: PMC4202328
- DOI: 10.1093/gbe/evu192
Natural selection on coding and noncoding DNA sequences is associated with virulence genes in a plant pathogenic fungus
Abstract
Natural selection leaves imprints on DNA, offering the opportunity to identify functionally important regions of the genome. Identifying the genomic regions affected by natural selection within pathogens can aid in the pursuit of effective strategies to control diseases. In this study, we analyzed genome-wide patterns of selection acting on different classes of sequences in a worldwide sample of eight strains of the model plant-pathogenic fungus Colletotrichum graminicola. We found evidence of selective sweeps, balancing selection, and positive selection affecting both protein-coding and noncoding DNA of pathogenicity-related sequences. Genes encoding putative effector proteins and secondary metabolite biosynthetic enzymes show evidence of positive selection acting on the coding sequence, consistent with an Arms Race model of evolution. The 5' untranslated regions (UTRs) of genes coding for effector proteins and genes upregulated during infection show an excess of high-frequency polymorphisms likely the consequence of balancing selection and consistent with the Red Queen hypothesis of evolution acting on these putative regulatory sequences. Based on the findings of this work, we propose that even though adaptive substitutions on coding sequences are important for proteins that interact directly with the host, polymorphisms in the regulatory sequences may confer flexibility of gene expression in the virulence processes of this important plant pathogen.
Keywords: Colletotrichum graminicola; PAML; arms race hypothesis; pathogenicity; positive selection.
© The Author(s) 2014. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures
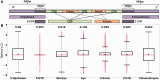

References
-
- Aguileta G, et al. Finding candidate genes under positive selection in non-model species: examples of genes involved in host specialization in pathogens. Mol Ecol. 2010;19:292–306. - PubMed
-
- Aguileta G, et al. Genes under positive selection in a model plant pathogenic fungus, Botrytis. Infect Genet Evol. 2012;12:987–996. - PubMed
-
- Andolfatto P. Adaptive evolution of non-coding DNA in Drosophila. Nature. 2005;437:1149–1152. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

