Critical roles of interdomain interactions for modulatory ATP binding to sarcoplasmic reticulum Ca2+-ATPase
- PMID: 25193668
- PMCID: PMC4200266
- DOI: 10.1074/jbc.M114.571687
Critical roles of interdomain interactions for modulatory ATP binding to sarcoplasmic reticulum Ca2+-ATPase
Abstract
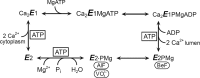
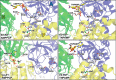
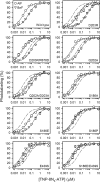
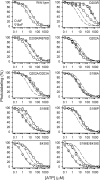
ATP has dual roles in the reaction cycle of sarcoplasmic reticulum Ca(2+)-ATPase. Upon binding to the Ca2E1 state, ATP phosphorylates the enzyme, and by binding to other conformational states in a non-phosphorylating modulatory mode ATP stimulates the dephosphorylation and other partial reaction steps of the cycle, thereby ensuring a high rate of Ca(2+) transport under physiological conditions. The present study elucidates the mechanism underlying the modulatory effect on dephosphorylation. In the intermediate states of dephosphorylation the A-domain residues Ser(186) and Asp(203) interact with Glu(439) (N-domain) and Arg(678) (P-domain), respectively. Single mutations to these residues abolish the stimulation of dephosphorylation by ATP. The double mutation swapping Asp(203) and Arg(678) rescues ATP stimulation, whereas this is not the case for the double mutation swapping Ser(186) and Glu(439). By taking advantage of the ability of wild type and mutant Ca(2+)-ATPases to form stable complexes with aluminum fluoride (E2·AlF) and beryllium fluoride (E2·BeF) as analogs of the E2·P phosphoryl transition state and E2P ground state, respectively, of the dephosphorylation reaction, the mutational effects on ATP binding to these intermediates are demonstrated. In the wild type Ca(2+)-ATPase, the ATP affinity of the E2·P phosphoryl transition state is higher than that of the E2P ground state, thus explaining the stimulation of dephosphorylation by nucleotide-induced transition state stabilization. We find that the Asp(203)-Arg(678) and Ser(186)-Glu(439) interdomain bonds are critical, because they tighten the interaction with ATP in the E2·P phosphoryl transition state. Moreover, ATP binding and the Ser(186)-Glu(439) bond are mutually exclusive in the E2P ground state.
Keywords: ATPase; Calcium ATPase; Enzyme Kinetics; Membrane Energetics; Membrane Enzyme; Membrane Transport; Phosphorylation; Site-directed Mutagenesis.
© 2014 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures


Similar articles
-
Modulatory ATP binding affinity in intermediate states of E2P dephosphorylation of sarcoplasmic reticulum Ca2+-ATPase.J Biol Chem. 2011 Apr 1;286(13):11792-802. doi: 10.1074/jbc.M110.206094. Epub 2011 Feb 2. J Biol Chem. 2011. PMID: 21288896 Free PMC article.
-
Distinct natures of beryllium fluoride-bound, aluminum fluoride-bound, and magnesium fluoride-bound stable analogues of an ADP-insensitive phosphoenzyme intermediate of sarcoplasmic reticulum Ca2+-ATPase: changes in catalytic and transport sites during phosphoenzyme hydrolysis.J Biol Chem. 2004 Apr 9;279(15):14991-8. doi: 10.1074/jbc.M313363200. Epub 2004 Jan 30. J Biol Chem. 2004. PMID: 14754887
-
Roles of conserved P domain residues and Mg2+ in ATP binding in the ground and Ca2+-activated states of sarcoplasmic reticulum Ca2+-ATPase.J Biol Chem. 2004 Jul 30;279(31):32515-23. doi: 10.1074/jbc.M403242200. Epub 2004 May 7. J Biol Chem. 2004. PMID: 15133025
-
Transport mechanism of the sarcoplasmic reticulum Ca2+ -ATPase pump.Curr Opin Struct Biol. 2005 Aug;15(4):387-93. doi: 10.1016/j.sbi.2005.06.005. Curr Opin Struct Biol. 2005. PMID: 16009548 Review.
-
Structural basis of the conformational and functional regulation of human SERCA2b, the ubiquitous endoplasmic reticulum calcium pump.Bioessays. 2022 Jul;44(7):e2200052. doi: 10.1002/bies.202200052. Epub 2022 May 13. Bioessays. 2022. PMID: 35560336 Review.
Cited by
-
Cryoelectron microscopy of Na+,K+-ATPase in the two E2P states with and without cardiotonic steroids.Proc Natl Acad Sci U S A. 2022 Apr 12;119(15):e2123226119. doi: 10.1073/pnas.2123226119. Epub 2022 Apr 5. Proc Natl Acad Sci U S A. 2022. PMID: 35380894 Free PMC article.
References
-
- Møller J. V., Olesen C., Winther A. M., Nissen P. (2010) The sarcoplasmic Ca2+-ATPase: design of a perfect chemi-osmotic pump. Q. Rev. Biophys. 43, 501–566 - PubMed
-
- Toyoshima C. (2009) How Ca2+-ATPase pumps ions across the sarcoplasmic reticulum membrane. Biochim. Biophys. Acta 1793, 941–946 - PubMed
-
- Toyoshima C., Iwasawa S., Ogawa H., Hirata A., Tsueda J., Inesi G. (2013) Crystal structures of the calcium pump and sarcolipin in the Mg2+-bound E1 state. Nature 495, 260–264 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

