Integrator regulates transcriptional initiation and pause release following activation
- PMID: 25201415
- PMCID: PMC4292851
- DOI: 10.1016/j.molcel.2014.08.004
Integrator regulates transcriptional initiation and pause release following activation
Abstract
In unicellular organisms, initiation is the rate-limiting step in transcription; in metazoan organisms, the transition from initiation to productive elongation is also important. Here, we show that the RNA polymerase II (RNAPII)-associated multiprotein complex, Integrator, plays a critical role in both initiation and the release of paused RNAPII at immediate early genes (IEGs) following transcriptional activation by epidermal growth factor (EGF) in human cells. Integrator is recruited to the IEGs in a signal-dependent manner and is required to engage and recruit the super elongation complex (SEC) to EGF-responsive genes to allow release of paused RNAPII and productive transcription elongation.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures



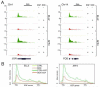
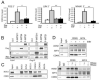
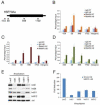

References
-
- Amit I, Citri A, Shay T, Lu Y, Katz M, Zhang F, Tarcic G, Siwak D, Lahad J, Jacob-Hirsch J, et al. A module of negative feedback regulators defines growth factor signaling. Nat Genet. 2007;39:503–512. - PubMed
-
- Baillat D, Hakimi MA, Naar AM, Shilatifard A, Cooch N, Shiekhattar R. Integrator, a multiprotein mediator of small nuclear RNA processing, associates with the C-terminal repeat of RNA polymerase II. Cell. 2005;123:265–276. - PubMed
-
- Barboric M, Nissen RM, Kanazawa S, Jabrane-Ferrat N, Peterlin BM. NF-kappaB binds P-TEFb to stimulate transcriptional elongation by RNA polymerase II. Mol Cell. 2001;8:327–337. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
