Asp-52 in combination with Asp-398 plays a critical role in ATP hydrolysis of chaperonin GroEL
- PMID: 25202010
- PMCID: PMC4208008
- DOI: 10.1074/jbc.M114.593822
Asp-52 in combination with Asp-398 plays a critical role in ATP hydrolysis of chaperonin GroEL
Abstract
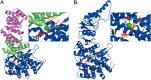
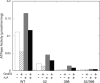
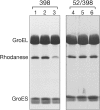
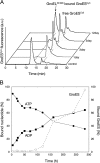
The Escherichia coli chaperonin GroEL is a double-ring chaperone that assists protein folding with the aid of GroES and ATP. Asp-398 in GroEL is known as one of the critical residues on ATP hydrolysis because GroEL(D398A) mutant is deficient in ATP hydrolysis (<2% of the wild type) but not in ATP binding. In the archaeal Group II chaperonin, another aspartate residue, Asp-52 in the corresponding E. coli GroEL, in addition to Asp-398 is also important for ATP hydrolysis. We investigated the role of Asp-52 in GroEL and found that ATPase activity of GroEL(D52A) and GroEL(D52A/D398A) mutants was ∼ 20% and <0.01% of wild-type GroEL, respectively, indicating that Asp-52 in E. coli GroEL is also involved in the ATP hydrolysis. GroEL(D52A/D398A) formed a symmetric football-shaped GroEL-GroES complex in the presence of ATP, again confirming the importance of the symmetric complex during the GroEL ATPase cycle. Notably, the symmetric complex of GroEL(D52A/D398A) was extremely stable, with a half-time of ∼ 150 h (∼ 6 days), providing a good model to characterize the football-shaped complex.
Keywords: ATPase; Chaperone; Chaperonin; GroEL; GroES; Protein Aggregation; Protein Folding; Protein Misfolding.
© 2014 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures


Similar articles
-
Revisiting the GroEL-GroES reaction cycle via the symmetric intermediate implied by novel aspects of the GroEL(D398A) mutant.J Biol Chem. 2008 Aug 29;283(35):23774-81. doi: 10.1074/jbc.M802542200. Epub 2008 Jun 20. J Biol Chem. 2008. PMID: 18567584 Free PMC article.
-
TEM and STEM-EDS evaluation of metal nanoparticle encapsulation in GroEL/GroES complexes according to the reaction mechanism of chaperonin.Microscopy (Oxf). 2021 Jun 6;70(3):289-296. doi: 10.1093/jmicro/dfaa064. Microscopy (Oxf). 2021. PMID: 33173948
-
Effective ATPase activity and moderate chaperonin-cochaperonin interaction are important for the functional single-ring chaperonin system.Biochem Biophys Res Commun. 2015 Oct 9;466(1):15-20. doi: 10.1016/j.bbrc.2015.08.034. Epub 2015 Aug 11. Biochem Biophys Res Commun. 2015. PMID: 26271593
-
Reaction Cycle of Chaperonin GroEL via Symmetric "Football" Intermediate.J Mol Biol. 2015 Sep 11;427(18):2912-8. doi: 10.1016/j.jmb.2015.04.007. Epub 2015 Apr 18. J Mol Biol. 2015. PMID: 25900372 Review.
-
GroEL and the GroEL-GroES Complex.Subcell Biochem. 2017;83:483-504. doi: 10.1007/978-3-319-46503-6_17. Subcell Biochem. 2017. PMID: 28271487 Review.
Cited by
-
Synergistic effects of ATP and RNA binding to human DEAD-box protein DDX1.Nucleic Acids Res. 2015 Mar 11;43(5):2813-28. doi: 10.1093/nar/gkv106. Epub 2015 Feb 17. Nucleic Acids Res. 2015. PMID: 25690890 Free PMC article.
-
Physicochemical Properties of the Mammalian Molecular Chaperone HSP60.Int J Mol Sci. 2018 Feb 6;19(2):489. doi: 10.3390/ijms19020489. Int J Mol Sci. 2018. PMID: 29415503 Free PMC article.
-
Pseudomonas aeruginosa GroEL Stimulates Production of PTX3 by Activating the NF-κB Pathway and Simultaneously Downregulating MicroRNA-9.Infect Immun. 2017 Feb 23;85(3):e00935-16. doi: 10.1128/IAI.00935-16. Print 2017 Mar. Infect Immun. 2017. PMID: 28031262 Free PMC article.
-
Chaperonin GroEL uses asymmetric and symmetric reaction cycles in response to the concentration of non-native substrate proteins.Biophys Physicobiol. 2016 Apr 22;13:63-69. doi: 10.2142/biophysico.13.0_63. eCollection 2016. Biophys Physicobiol. 2016. PMID: 27924258 Free PMC article. Review.
-
Pathogenic mutation impairs functional dynamics of Hsp60 in mono- and oligomeric states.Nat Commun. 2025 Apr 3;16(1):3158. doi: 10.1038/s41467-025-57958-5. Nat Commun. 2025. PMID: 40180932 Free PMC article.
References
-
- Richter K., Haslbeck M., Buchner J. (2010) The heat shock response: life on the verge of death. Mol. Cell 40, 253–266 - PubMed
-
- Yébenes H., Mesa P., Muñoz I. G., Montoya G., Valpuesta J. M. (2011) Chaperonins: two rings for folding. Trends Biochem. Sci. 36, 424–432 - PubMed
-
- Saibil H. R., Fenton W. A., Clare D. K., Horwich A. L. (2013) Structure and allostery of the chaperonin GroEL. J. Mol. Biol. 425, 1476–1487 - PubMed
-
- Kim Y. E., Hipp M. S., Bracher A., Hayer-Hartl M., Hartl F. U. (2013) Molecular chaperone functions in protein folding and proteostasis. Annu. Rev. Biochem. 82, 323–355 - PubMed
-
- Braig K., Otwinowski Z., Hegde R., Boisvert D. C., Joachimiak A., Horwich A. L., Sigler P. B. (1994) The crystal structure of the bacterial chaperonin GroEL at 2.8 Å. Nature 371, 578–586 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

