A comparative metagenome survey of the fecal microbiota of a breast- and a plant-fed Asian elephant reveals an unexpectedly high diversity of glycoside hydrolase family enzymes
- PMID: 25208077
- PMCID: PMC4160196
- DOI: 10.1371/journal.pone.0106707
A comparative metagenome survey of the fecal microbiota of a breast- and a plant-fed Asian elephant reveals an unexpectedly high diversity of glycoside hydrolase family enzymes
Abstract
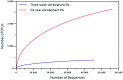
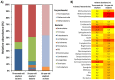
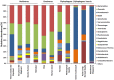
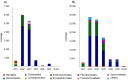
A phylogenetic and metagenomic study of elephant feces samples (derived from a three-weeks-old and a six-years-old Asian elephant) was conducted in order to describe the microbiota inhabiting this large land-living animal. The microbial diversity was examined via 16S rRNA gene analysis. We generated more than 44,000 GS-FLX+454 reads for each animal. For the baby elephant, 380 operational taxonomic units (OTUs) were identified at 97% sequence identity level; in the six-years-old animal, close to 3,000 OTUs were identified, suggesting high microbial diversity in the older animal. In both animals most OTUs belonged to Bacteroidetes and Firmicutes. Additionally, for the baby elephant a high number of Proteobacteria was detected. A metagenomic sequencing approach using Illumina technology resulted in the generation of 1.1 Gbp assembled DNA in contigs with a maximum size of 0.6 Mbp. A KEGG pathway analysis suggested high metabolic diversity regarding the use of polymers and aromatic and non-aromatic compounds. In line with the high phylogenetic diversity, a surprising and not previously described biodiversity of glycoside hydrolase (GH) genes was found. Enzymes of 84 GH families were detected. Polysaccharide utilization loci (PULs), which are found in Bacteroidetes, were highly abundant in the dataset; some of these comprised cellulase genes. Furthermore the highest coverage for GH5 and GH9 family enzymes was detected for Bacteroidetes, suggesting that bacteria of this phylum are mainly responsible for the degradation of cellulose in the Asian elephant. Altogether, this study delivers insight into the biomass conversion by one of the largest plant-fed and land-living animals.
Conflict of interest statement
Figures

References
-
- Björnhag G (1994) The Digestive System in Mammals: Food, Form and Function: Cambridge University Press: 287–309.
-
- Flint HJ, Bayer EA, Rincon MT, Lamed R, White BA (2008) Polysaccharide utilization by gut bacteria: potential for new insights from genomic analysis. Nature Reviews Microbiology 6: 121–131. - PubMed
-
- Bayané A, Guiot S (2011) Animal digestive strategies versus anaerobic digestion bioprocesses for biogas production from lignocellulosic biomass. Reviews in Environmental Science and Bio/Technology 10: 43–62.
-
- Clauss M, Frey R, Kiefer B, Lechner-Doll M, Loehlein W, et al. (2003) The maximum attainable body size of herbivorous mammals: morphophysiological constraints on foregut, and adaptations of hindgut fermenters. Oecologia 136: 14–27. - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

