Architecture of mammalian respiratory complex I
- PMID: 25209663
- PMCID: PMC4224586
- DOI: 10.1038/nature13686
Architecture of mammalian respiratory complex I
Abstract
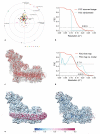
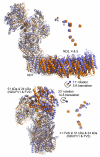
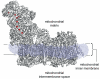
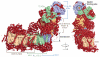
Complex I (NADH:ubiquinone oxidoreductase) is essential for oxidative phosphorylation in mammalian mitochondria. It couples electron transfer from NADH to ubiquinone with proton translocation across the energy-transducing inner membrane, providing electrons for respiration and driving ATP synthesis. Mammalian complex I contains 44 different nuclear- and mitochondrial-encoded subunits, with a combined mass of 1 MDa. The 14 conserved 'core' subunits have been structurally defined in the minimal, bacterial complex, but the structures and arrangement of the 30 'supernumerary' subunits are unknown. Here we describe a 5 Å resolution structure of complex I from Bos taurus heart mitochondria, a close relative of the human enzyme, determined by single-particle electron cryo-microscopy. We present the structures of the mammalian core subunits that contain eight iron-sulphur clusters and 60 transmembrane helices, identify 18 supernumerary transmembrane helices, and assign and model 14 supernumerary subunits. Thus, we considerably advance knowledge of the structure of mammalian complex I and the architecture of its supernumerary ensemble around the core domains. Our structure provides insights into the roles of the supernumerary subunits in regulation, assembly and homeostasis, and a basis for understanding the effects of mutations that cause a diverse range of human diseases.
Figures






Similar articles
-
Structure of mammalian respiratory complex I.Nature. 2016 Aug 18;536(7616):354-358. doi: 10.1038/nature19095. Epub 2016 Aug 10. Nature. 2016. PMID: 27509854 Free PMC article.
-
Structure of subcomplex Iβ of mammalian respiratory complex I leads to new supernumerary subunit assignments.Proc Natl Acad Sci U S A. 2015 Sep 29;112(39):12087-92. doi: 10.1073/pnas.1510577112. Epub 2015 Sep 14. Proc Natl Acad Sci U S A. 2015. PMID: 26371297 Free PMC article.
-
Atomic structure of the entire mammalian mitochondrial complex I.Nature. 2016 Oct 20;538(7625):406-410. doi: 10.1038/nature19794. Epub 2016 Sep 5. Nature. 2016. PMID: 27595392 Free PMC article.
-
The nuclear encoded subunits of complex I from bovine heart mitochondria.Biochim Biophys Acta. 2003 Jul 10;1604(3):135-50. doi: 10.1016/s0005-2728(03)00059-8. Biochim Biophys Acta. 2003. PMID: 12837546 Review.
-
Mammalian Mitochondrial Complex I Structure and Disease-Causing Mutations.Trends Cell Biol. 2018 Oct;28(10):835-867. doi: 10.1016/j.tcb.2018.06.006. Epub 2018 Jul 26. Trends Cell Biol. 2018. PMID: 30055843 Review.
Cited by
-
Mechanisms of Mitochondrial ROS Production in Assisted Reproduction: The Known, the Unknown, and the Intriguing.Antioxidants (Basel). 2020 Sep 29;9(10):933. doi: 10.3390/antiox9100933. Antioxidants (Basel). 2020. PMID: 33003362 Free PMC article. Review.
-
Mitochondrial Stress in Metabolic Inflammation: Modest Benefits and Full Losses.Oxid Med Cell Longev. 2022 Nov 22;2022:8803404. doi: 10.1155/2022/8803404. eCollection 2022. Oxid Med Cell Longev. 2022. PMID: 36457729 Free PMC article. Review.
-
The Role of Metformin in Controlling Oxidative Stress in Muscle of Diabetic Rats.Oxid Med Cell Longev. 2016;2016:6978625. doi: 10.1155/2016/6978625. Epub 2016 Aug 4. Oxid Med Cell Longev. 2016. PMID: 27579154 Free PMC article.
-
Creation of Mitochondrial Disease Models Using Mitochondrial DNA Editing.Biomedicines. 2023 Feb 12;11(2):532. doi: 10.3390/biomedicines11020532. Biomedicines. 2023. PMID: 36831068 Free PMC article. Review.
-
NDUFAB1 confers cardio-protection by enhancing mitochondrial bioenergetics through coordination of respiratory complex and supercomplex assembly.Cell Res. 2019 Sep;29(9):754-766. doi: 10.1038/s41422-019-0208-x. Epub 2019 Jul 31. Cell Res. 2019. PMID: 31366990 Free PMC article.
References
-
- Hirst J. Mitochondrial complex I. Annu. Rev. Biochem. 2013;82:551–575. - PubMed
-
- Carroll J, Fearnley IM, Shannon RJ, Hirst J, Walker JE. Analysis of the subunit composition of complex I from bovine heart mitochondria. Mol. Cell. Proteomics. 2003;2:117–126. - PubMed
-
- Hirst J, Carroll J, Fearnley IM, Shannon RJ, Walker JE. The nuclear encoded subunits of complex I from bovine heart mitochondria. Biochim. Biophys. Acta. 2003;1604:135–150. - PubMed
-
- Efremov RG, Sazanov LA. Structure of the membrane domain of respiratory complex I. Nature. 2011;476:414–420. - PubMed
ADDITIONAL METHODS REFERENCES
-
- Bellare JR, Davis HT, Scriven LE, Talmon Y. Controlled environment vitrification system: an improved sample preparation technique. J. Electron Microsc. Tech. 1988;10:87–111. - PubMed
-
- Grigorieff N. FREALIGN: high-resolution refinement of single particle structures. J. Struct. Biol. 2007;157:117–125. - PubMed
-
- Smith JM. XIMDISP – a visualization tool to aid structure determination from electron microscope images. J. Struct. Biol. 1999;125:223–228. - PubMed
-
- Tang G, et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 2007;157:38–46. - PubMed
ADDITIONAL EXTENDED DATA REFERENCES
-
- Efremov RG, Sazanov LA. Respiratory complex I: ‘steam engine’ of the cell? Curr. Opin. Struct. Biol. 2011;21:532–540. - PubMed
-
- Krissinel E, Henrick K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Cryst. 2004;D60:2256–2268. - PubMed
-
- Balsa E, et al. NDUFA4 is a subunit of complex IV of the mammalian electron transport chain. Cell Metab. 2012;16:378–386. - PubMed
-
- Johansson K, et al. Structural basis for substrate specificities of cellular deoxyribonucleoside kinases. Nat. Struct. Biol. 2001;8:616–620. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

