Urzymology: experimental access to a key transition in the appearance of enzymes
- PMID: 25210034
- PMCID: PMC4215205
- DOI: 10.1074/jbc.R114.567495
Urzymology: experimental access to a key transition in the appearance of enzymes
Abstract
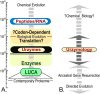
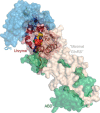
Urzymes are catalysts derived from invariant cores of protein superfamilies. Urzymes from both aminoacyl-tRNA synthetase classes possess sophisticated catalytic mechanisms: pre-steady state bursts, significant transition-state stabilization of both amino acid activation, and tRNA acylation. However, they have insufficient specificity to ensure a fully developed genetic code, suggesting that they participated in synthesizing statistical proteins. They represent a robust experimental platform from which to articulate and test hypotheses both about their own ancestors and about how they, in turn, evolved into modern enzymes. They help reshape numerous paradigms from the RNA World hypothesis to protein structure databases and allostery.
Keywords: Aminoacyl tRNA Synthetase; Bioinformatics; Genetic Code; Precellular Evolution; Protein Engineering; Protein Evolution; Protein Synthesis.
© 2014 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures

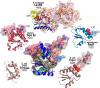
Similar articles
-
The Rodin-Ohno hypothesis that two enzyme superfamilies descended from one ancestral gene: an unlikely scenario for the origins of translation that will not be dismissed.Biol Direct. 2014 Jun 14;9:11. doi: 10.1186/1745-6150-9-11. Biol Direct. 2014. PMID: 24927791 Free PMC article.
-
Coding of Class I and II Aminoacyl-tRNA Synthetases.Adv Exp Med Biol. 2017;966:103-148. doi: 10.1007/5584_2017_93. Adv Exp Med Biol. 2017. PMID: 28828732 Free PMC article. Review.
-
Aminoacylating urzymes challenge the RNA world hypothesis.J Biol Chem. 2013 Sep 13;288(37):26856-63. doi: 10.1074/jbc.M113.496125. Epub 2013 Jul 18. J Biol Chem. 2013. PMID: 23867455 Free PMC article.
-
Structural phylogenomics retrodicts the origin of the genetic code and uncovers the evolutionary impact of protein flexibility.PLoS One. 2013 Aug 21;8(8):e72225. doi: 10.1371/journal.pone.0072225. eCollection 2013. PLoS One. 2013. PMID: 23991065 Free PMC article.
-
Structure, function and evolution of seryl-tRNA synthetases: implications for the evolution of aminoacyl-tRNA synthetases and the genetic code.J Mol Evol. 1995 May;40(5):519-30. doi: 10.1007/BF00166620. J Mol Evol. 1995. PMID: 7540217 Review.
Cited by
-
11th IUBMB Focused Meeting on the Aminoacyl-tRNA Synthetases: Sailing a New Sea of Complex Functions in Human Biology and Disease.Biomolecules. 2018 May 1;8(2):22. doi: 10.3390/biom8020022. Biomolecules. 2018. PMID: 29723968 Free PMC article.
-
Recapitulating the Structural Evolution of Redox Regulation in Adenosine 5'-Phosphosulfate Kinase from Cyanobacteria to Plants.J Biol Chem. 2015 Oct 9;290(41):24705-14. doi: 10.1074/jbc.M115.679514. Epub 2015 Aug 20. J Biol Chem. 2015. PMID: 26294763 Free PMC article.
-
Evolution of the genetic code.Transcription. 2021 Feb;12(1):28-53. doi: 10.1080/21541264.2021.1927652. Epub 2021 May 18. Transcription. 2021. PMID: 34000965 Free PMC article. Review.
-
Mycobacterium tuberculosis Phe-tRNA synthetase: structural insights into tRNA recognition and aminoacylation.Nucleic Acids Res. 2021 May 21;49(9):5351-5368. doi: 10.1093/nar/gkab272. Nucleic Acids Res. 2021. PMID: 33885823 Free PMC article.
-
Impedance Matching and the Choice Between Alternative Pathways for the Origin of Genetic Coding.Int J Mol Sci. 2020 Oct 7;21(19):7392. doi: 10.3390/ijms21197392. Int J Mol Sci. 2020. PMID: 33036401 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

