Genetic analysis of the ribosome biogenesis factor Ltv1 of Saccharomyces cerevisiae
- PMID: 25213169
- PMCID: PMC4224153
- DOI: 10.1534/genetics.114.168294
Genetic analysis of the ribosome biogenesis factor Ltv1 of Saccharomyces cerevisiae
Abstract
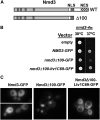
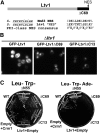
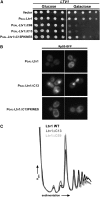
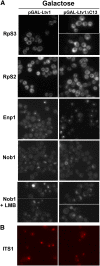
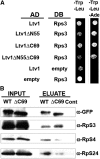
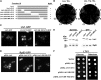
Ribosome biogenesis has been studied extensively in the yeast Saccharomyces cerevisiae. Yeast Ltv1 is a conserved 40S-associated biogenesis factor that has been proposed to function in small subunit nuclear export. Here we show that Ltv1 has a canonical leucine-rich nuclear export signal (NES) at its extreme C terminus that is both necessary for Crm1 interaction and Ltv1 export. The C terminus of Ltv1 can substitute for the NES in the 60S-export adapter Nmd3, demonstrating that it is a functional NES. Overexpression of an Ltv1 lacking its NES (Ltv1∆C13) was strongly dominant negative and resulted in the nuclear accumulation of RpS3-GFP; however, export of the pre-40S was not affected. In addition, expression of endogenous levels of Ltv1∆C protein complemented both the slow-growth phenotype and the 40S biogenesis defect of an ltv1 deletion mutant. Thus, if Ltv1 is a nuclear export adapter for the pre-40S subunit, its function must be fully redundant with additional export factors. The dominant negative phenotype of Ltv1∆NES overexpression was suppressed by co-overexpressing RpS3 and its chaperone, Yar1, or by deletion of the RpS3-binding site in Ltv1∆NES, suggesting that titration of RpS3 by Ltv1∆NES is deleterious in yeast. The dominant-negative phenotype did not correlate with a decrease in 40S levels but rather with a reduction in the polysome-to-monosome ratio, indicating reduced rates of translation. We suggest that titration of RpS3 by excess nuclear Ltv1 interferes with 40S function or with a nonribosomal function of RpS3.
Keywords: Ltv1; RpS3; nuclear export; ribosome biogenesis; small subunit.
Copyright © 2014 by the Genetics Society of America.
Figures



Comment in
-
Location is everything: an educational primer for use with "genetic analysis of the ribosome biogenesis factor Ltv1 of Saccharomyces cerevisiae".Genetics. 2015 Feb;199(2):307-13. doi: 10.1534/genetics.114.173641. Genetics. 2015. PMID: 25657348 Free PMC article.
Similar articles
-
Dominant mutations in the late 40S biogenesis factor Ltv1 affect cytoplasmic maturation of the small ribosomal subunit in Saccharomyces cerevisiae.Genetics. 2010 May;185(1):199-209. doi: 10.1534/genetics.110.115584. Epub 2010 Mar 9. Genetics. 2010. PMID: 20215468 Free PMC article.
-
Ltv1 is required for efficient nuclear export of the ribosomal small subunit in Saccharomyces cerevisiae.Genetics. 2006 Oct;174(2):679-91. doi: 10.1534/genetics.106.062117. Epub 2006 Aug 3. Genetics. 2006. PMID: 16888326 Free PMC article.
-
Genetic and biochemical interactions among Yar1, Ltv1 and Rps3 define novel links between environmental stress and ribosome biogenesis in Saccharomyces cerevisiae.Genetics. 2004 Dec;168(4):1877-89. doi: 10.1534/genetics.104.032656. Genetics. 2004. PMID: 15611164 Free PMC article.
-
Nuclear export and cytoplasmic maturation of ribosomal subunits.FEBS Lett. 2007 Jun 19;581(15):2783-93. doi: 10.1016/j.febslet.2007.05.013. Epub 2007 May 11. FEBS Lett. 2007. PMID: 17509569 Review.
-
Nuclear export of ribosomal subunits.Trends Biochem Sci. 2002 Nov;27(11):580-5. doi: 10.1016/s0968-0004(02)02208-9. Trends Biochem Sci. 2002. PMID: 12417134 Review.
Cited by
-
Sequential domain assembly of ribosomal protein S3 drives 40S subunit maturation.Nat Commun. 2016 Feb 2;7:10336. doi: 10.1038/ncomms10336. Nat Commun. 2016. PMID: 26831757 Free PMC article.
-
Genetic dissection of early endosomal recycling highlights a TORC1-independent role for Rag GTPases.J Cell Biol. 2017 Oct 2;216(10):3275-3290. doi: 10.1083/jcb.201702177. Epub 2017 Aug 2. J Cell Biol. 2017. PMID: 28768685 Free PMC article.
-
A non-canonical mechanism for Crm1-export cargo complex assembly.Elife. 2015 Apr 21;4:e05745. doi: 10.7554/eLife.05745. Elife. 2015. PMID: 25895666 Free PMC article.
-
DNA methylation is associated with lung function in never smokers.Respir Res. 2019 Dec 2;20(1):268. doi: 10.1186/s12931-019-1222-8. Respir Res. 2019. PMID: 31791327 Free PMC article.
-
Drosophila Low Temperature Viability Protein 1 (LTV1) Is Required for Ribosome Biogenesis and Cell Growth Downstream of Drosophila Myc (dMyc).J Biol Chem. 2015 May 22;290(21):13591-604. doi: 10.1074/jbc.M114.607036. Epub 2015 Apr 9. J Biol Chem. 2015. PMID: 25858587 Free PMC article.
References
-
- Amberg D. C., Goldstein A. L., Cole C. N., 1992. Isolation and characterization of RAT1: an essential gene of Saccharomyces cerevisiae required for the efficient nucleocytoplasmic trafficking of mRNA. Genes Dev. 6: 1173–1189. - PubMed
-
- Bassler J., Grandi P., Gadal O., Lessmann T., Petfalski E., et al. , 2001. Identification of a 60S preribosomal particle that is closely linked to nuclear export. Mol. Cell 8: 517–529. - PubMed
-
- Bradatsch B., Katahira J., Kowalinski E., Bange G., Yao W., et al. , 2007. Arx1 functions as an unorthodox nuclear export receptor for the 60S preribosomal subunit. Mol. Cell 27: 767–779. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

