Arabidopsis thaliana RNase H2 deficiency counteracts the needs for the WEE1 checkpoint kinase but triggers genome instability
- PMID: 25217508
- PMCID: PMC4213155
- DOI: 10.1105/tpc.114.128108
Arabidopsis thaliana RNase H2 deficiency counteracts the needs for the WEE1 checkpoint kinase but triggers genome instability
Abstract
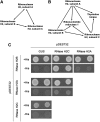
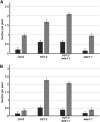
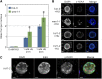
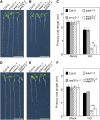
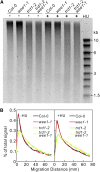
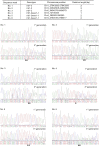
The WEE1 kinase is an essential cell cycle checkpoint regulator in Arabidopsis thaliana plants experiencing replication defects. Whereas under non-stress conditions WEE1-deficient plants develop normally, they fail to adapt to replication inhibitory conditions, resulting in the accumulation of DNA damage and loss of cell division competence. We identified mutant alleles of the genes encoding subunits of the ribonuclease H2 (RNase H2) complex, known for its role in removing ribonucleotides from DNA-RNA duplexes, as suppressor mutants of WEE1 knockout plants. RNase H2 deficiency triggered an increase in homologous recombination (HR), correlated with the accumulation of γ-H2AX foci. However, as HR negatively impacts the growth of WEE1-deficient plants under replication stress, it cannot account for the rescue of the replication defects of the WEE1 knockout plants. Rather, the observed increase in ribonucleotide incorporation in DNA indicates that the substitution of deoxynucleotide with ribonucleotide abolishes the need for WEE1 under replication stress. Strikingly, increased ribonucleotide incorporation in DNA correlated with the occurrence of small base pair deletions, identifying the RNase H2 complex as an important suppressor of genome instability.
© 2014 American Society of Plant Biologists. All rights reserved.
Figures


Similar articles
-
Lack of RNase H2 activity rescues HU-sensitivity of WEE1 deficient plants.Plant Signal Behav. 2015;10(4):e1001226. doi: 10.1080/15592324.2014.1001226. Plant Signal Behav. 2015. PMID: 25875879 Free PMC article.
-
The Arabidopsis thaliana checkpoint kinase WEE1 protects against premature vascular differentiation during replication stress.Plant Cell. 2011 Apr;23(4):1435-48. doi: 10.1105/tpc.110.082768. Epub 2011 Apr 15. Plant Cell. 2011. PMID: 21498679 Free PMC article.
-
A Mutation in DNA Polymerase α Rescues WEE1KO Sensitivity to HU.Int J Mol Sci. 2021 Aug 30;22(17):9409. doi: 10.3390/ijms22179409. Int J Mol Sci. 2021. PMID: 34502313 Free PMC article.
-
Aicardi-Goutières syndrome: clues from the RNase H2 knock-out mouse.J Mol Med (Berl). 2013 Nov;91(11):1235-40. doi: 10.1007/s00109-013-1061-x. Epub 2013 Jun 7. J Mol Med (Berl). 2013. PMID: 23744109 Review.
-
Molecular and physiological consequences of faulty eukaryotic ribonucleotide excision repair.EMBO J. 2020 Feb 3;39(3):e102309. doi: 10.15252/embj.2019102309. Epub 2019 Dec 12. EMBO J. 2020. PMID: 31833079 Free PMC article. Review.
Cited by
-
Distinct features of ribonucleotides within genomic DNA in Aicardi-Goutières syndrome (AGS)-ortholog mutants of Saccharomyces cerevisiae.bioRxiv [Preprint]. 2023 Oct 2:2023.10.02.560505. doi: 10.1101/2023.10.02.560505. bioRxiv. 2023. Update in: iScience. 2024 May 16;27(6):110012. doi: 10.1016/j.isci.2024.110012. PMID: 37873120 Free PMC article. Updated. Preprint.
-
G2/M-checkpoint activation in fasciata1 rescues an aberrant S-phase checkpoint but causes genome instability.Plant Physiol. 2021 Aug 3;186(4):1893-1907. doi: 10.1093/plphys/kiab201. Plant Physiol. 2021. PMID: 34618100 Free PMC article.
-
The ATR-WEE1 kinase module inhibits the MAC complex to regulate replication stress response.Nucleic Acids Res. 2021 Feb 22;49(3):1411-1425. doi: 10.1093/nar/gkaa1082. Nucleic Acids Res. 2021. PMID: 33450002 Free PMC article.
-
Disproportionate presence of adenosine in mitochondrial and chloroplast DNA of Chlamydomonas reinhardtii.iScience. 2020 Dec 29;24(1):102005. doi: 10.1016/j.isci.2020.102005. eCollection 2021 Jan 22. iScience. 2020. PMID: 33490913 Free PMC article.
-
Buying time: FASCIATA1 deficiency rescues wee1 plants from replication stress by delaying mitosis.Plant Physiol. 2021 Aug 3;186(4):1762-1764. doi: 10.1093/plphys/kiab256. Plant Physiol. 2021. PMID: 34618115 Free PMC article. No abstract available.
References
-
- Bleuyard, J.Y., Gallego, M.E., Savigny, F., White, C.I. (2005). Differing requirements for the Arabidopsis Rad51 paralogs in meiosis and DNA repair. Plant J. 41: 533–545 - PubMed
-
- Boudolf, V., Inzé, D., De Veylder, L. (2006). What if higher plants lack a CDC25 phosphatase? Trends Plant Sci. 11: 474–479 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

