A gemcitabine sensitivity screen identifies a role for NEK9 in the replication stress response
- PMID: 25217585
- PMCID: PMC4191414
- DOI: 10.1093/nar/gku840
A gemcitabine sensitivity screen identifies a role for NEK9 in the replication stress response
Abstract
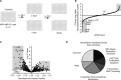
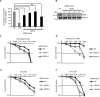
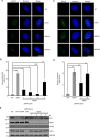
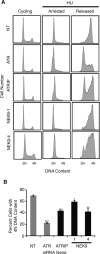
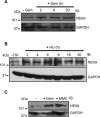
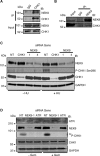
The Replication Stress Response (RSR) is a signaling network that recognizes challenges to DNA replication and coordinates diverse DNA repair and cell-cycle checkpoint pathways. Gemcitabine is a nucleoside analogue that causes cytotoxicity by inducing DNA replication blocks. Using a synthetic lethal screen of a RNAi library of nuclear enzymes to identify genes that when silenced cause gemcitabine sensitization or resistance in human triple-negative breast cancer cells, we identified NIMA (never in mitosis gene A)-related kinase 9 (NEK9) as a key component of the RSR. NEK9 depletion in cells leads to replication stress hypersensitivity, spontaneous accumulation of DNA damage and RPA70 foci, and an impairment in recovery from replication arrest. NEK9 protein levels also increase in response to replication stress. NEK9 complexes with CHK1, and moreover, NEK9 depletion impairs CHK1 autophosphorylation and kinase activity in response to replication stress. Thus, NEK9 is a critical component of the RSR that promotes CHK1 activity, maintaining genome integrity following challenges to DNA replication.
© The Author(s) 2014. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
Similar articles
-
Damage-induced DNA replication stalling relies on MAPK-activated protein kinase 2 activity.Proc Natl Acad Sci U S A. 2013 Oct 15;110(42):16856-61. doi: 10.1073/pnas.1304355110. Epub 2013 Sep 30. Proc Natl Acad Sci U S A. 2013. PMID: 24082115 Free PMC article.
-
H2AX phosphorylation marks gemcitabine-induced stalled replication forks and their collapse upon S-phase checkpoint abrogation.Mol Cancer Ther. 2007 Apr;6(4):1239-48. doi: 10.1158/1535-7163.MCT-06-0633. Epub 2007 Apr 3. Mol Cancer Ther. 2007. PMID: 17406032
-
The MAPK-activated protein kinase 2 mediates gemcitabine sensitivity in pancreatic cancer cells.Cell Cycle. 2014;13(6):884-9. doi: 10.4161/cc.28292. Epub 2014 Feb 21. Cell Cycle. 2014. PMID: 24556918 Free PMC article.
-
The ATM-Chk2 and ATR-Chk1 pathways in DNA damage signaling and cancer.Adv Cancer Res. 2010;108:73-112. doi: 10.1016/B978-0-12-380888-2.00003-0. Adv Cancer Res. 2010. PMID: 21034966 Review.
-
Safeguarding genome integrity: the checkpoint kinases ATR, CHK1 and WEE1 restrain CDK activity during normal DNA replication.Nucleic Acids Res. 2012 Jan;40(2):477-86. doi: 10.1093/nar/gkr697. Epub 2011 Sep 21. Nucleic Acids Res. 2012. PMID: 21937510 Free PMC article. Review.
Cited by
-
Cytidine deaminases APOBEC3C and APOBEC3D promote DNA replication stress resistance in pancreatic cancer cells.Nat Cancer. 2024 Jun;5(6):895-915. doi: 10.1038/s43018-024-00742-z. Epub 2024 Mar 6. Nat Cancer. 2024. PMID: 38448522
-
Activator protein 1 promotes gemcitabine-induced apoptosis in pancreatic cancer by upregulating its downstream target Bim.Oncol Lett. 2016 Dec;12(6):4732-4738. doi: 10.3892/ol.2016.5294. Epub 2016 Oct 19. Oncol Lett. 2016. PMID: 28105181 Free PMC article.
-
Overgrazing induces alterations in the hepatic proteome of sheep (Ovis aries): an iTRAQ-based quantitative proteomic analysis.Proteome Sci. 2017 Jan 5;15:2. doi: 10.1186/s12953-016-0111-z. eCollection 2016. Proteome Sci. 2017. PMID: 28149202 Free PMC article.
-
mTORC2 regulates ribonucleotide reductase to promote DNA replication and gemcitabine resistance in non-small cell lung cancer.Neoplasia. 2021 Jul;23(7):643-652. doi: 10.1016/j.neo.2021.05.007. Epub 2021 Jun 11. Neoplasia. 2021. PMID: 34126361 Free PMC article.
-
Hacking the Cell: Network Intrusion and Exploitation by Adenovirus E1A.mBio. 2018 May 1;9(3):e00390-18. doi: 10.1128/mBio.00390-18. mBio. 2018. PMID: 29717008 Free PMC article. Review.
References
-
- Zou L., Elledge S.J. Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science. 2003;300:1542–1548. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

