Hyb-Seq: Combining target enrichment and genome skimming for plant phylogenomics
- PMID: 25225629
- PMCID: PMC4162667
- DOI: 10.3732/apps.1400042
Hyb-Seq: Combining target enrichment and genome skimming for plant phylogenomics
Abstract
Premise of the study: Hyb-Seq, the combination of target enrichment and genome skimming, allows simultaneous data collection for low-copy nuclear genes and high-copy genomic targets for plant systematics and evolution studies. •
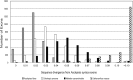
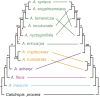
Methods and results: Genome and transcriptome assemblies for milkweed (Asclepias syriaca) were used to design enrichment probes for 3385 exons from 768 genes (>1.6 Mbp) followed by Illumina sequencing of enriched libraries. Hyb-Seq of 12 individuals (10 Asclepias species and two related genera) resulted in at least partial assembly of 92.6% of exons and 99.7% of genes and an average assembly length >2 Mbp. Importantly, complete plastomes and nuclear ribosomal DNA cistrons were assembled using off-target reads. Phylogenomic analyses demonstrated signal conflict between genomes. •
Conclusions: The Hyb-Seq approach enables targeted sequencing of thousands of low-copy nuclear exons and flanking regions, as well as genome skimming of high-copy repeats and organellar genomes, to efficiently produce genome-scale data sets for phylogenomics.
Keywords: Hyb-Seq; genome skimming; nuclear loci; phylogenomics; species tree; target enrichment.
Figures
Similar articles
-
Navigating the tip of the genomic iceberg: Next-generation sequencing for plant systematics.Am J Bot. 2012 Feb;99(2):349-64. doi: 10.3732/ajb.1100335. Epub 2011 Dec 14. Am J Bot. 2012. PMID: 22174336
-
Developing Asparagaceae1726: An Asparagaceae-specific probe set targeting 1726 loci for Hyb-Seq and phylogenomics in the family.Appl Plant Sci. 2024 Jun 18;12(5):e11597. doi: 10.1002/aps3.11597. eCollection 2024 Sep-Oct. Appl Plant Sci. 2024. PMID: 39360194 Free PMC article.
-
A pipeline for assembling low copy nuclear markers from plant genome skimming data for phylogenetic use.PeerJ. 2022 Dec 6;10:e14525. doi: 10.7717/peerj.14525. eCollection 2022. PeerJ. 2022. PMID: 36523475 Free PMC article.
-
Plant phylogenomics based on genome-partitioning strategies: Progress and prospects.Plant Divers. 2018 Jun 30;40(4):158-164. doi: 10.1016/j.pld.2018.06.005. eCollection 2018 Aug. Plant Divers. 2018. PMID: 30740560 Free PMC article. Review.
-
Practical considerations for plant phylogenomics.Appl Plant Sci. 2018 Apr 2;6(3):e1038. doi: 10.1002/aps3.1038. eCollection 2018 Mar. Appl Plant Sci. 2018. PMID: 29732268 Free PMC article. Review.
Cited by
-
Phylogenomic insights into Neotropical Magnolia relationships.Heliyon. 2024 Oct 16;10(20):e39430. doi: 10.1016/j.heliyon.2024.e39430. eCollection 2024 Oct 30. Heliyon. 2024. PMID: 39469672 Free PMC article.
-
Exploring Taxonomic and Genetic Relationships in the Pinus mugo Complex Using Genome Skimming Data.Int J Mol Sci. 2024 Sep 22;25(18):10178. doi: 10.3390/ijms251810178. Int J Mol Sci. 2024. PMID: 39337663 Free PMC article.
-
A Universal Probe Set for Targeted Sequencing of 353 Nuclear Genes from Any Flowering Plant Designed Using k-Medoids Clustering.Syst Biol. 2019 Jul 1;68(4):594-606. doi: 10.1093/sysbio/syy086. Syst Biol. 2019. PMID: 30535394 Free PMC article.
-
Capturing variation in Lens (Fabaceae): Development and utility of an exome capture array for lentil.Appl Plant Sci. 2018 Jul 16;6(7):e01165. doi: 10.1002/aps3.1165. eCollection 2018 Jun. Appl Plant Sci. 2018. PMID: 30131907 Free PMC article.
-
Phylogenomic analyses of the East Asian endemic Abelia (Caprifoliaceae) shed insights into the temporal and spatial diversification history with widespread hybridization.Ann Bot. 2022 Jan 28;129(2):201-216. doi: 10.1093/aob/mcab139. Ann Bot. 2022. PMID: 34950959 Free PMC article.
References
-
- Chapman M. A., Chang J., Weisman D., Kesseli R. V., Burke J. M. 2007. Universal markers for comparative mapping and phylogenetic analysis in the Asteraceae (Compositae). Theoretical and Applied Genetics 115: 747–755 - PubMed
-
- Cronn R., Knaus B. J., Liston A., Maughan P. J., Parks M., Syring J. V., Udall J. 2012. Targeted enrichment strategies for next-generation plant biology. American Journal of Botany 99: 291–311 - PubMed
-
- Davey J. W., Hohenlohe P. A., Etter P. D., Boone J. Q., Catchen J. M., Blaxter M. L. 2011. Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nature Reviews. Genetics 12: 499–510 - PubMed
-
- Doyle J. J., Doyle J. L. 1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 19: 11–15
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

