Adiposity significantly modifies genetic risk for dyslipidemia
- PMID: 25225679
- PMCID: PMC4617143
- DOI: 10.1194/jlr.P052522
Adiposity significantly modifies genetic risk for dyslipidemia
Abstract
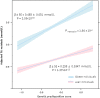
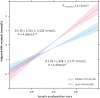
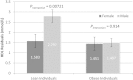
Recent genome-wide association studies have identified multiple loci robustly associated with plasma lipids, which also contribute to extreme lipid phenotypes. However, these common genetic variants explain <12% of variation in lipid traits. Adiposity is also an important determinant of plasma lipoproteins, particularly plasma TGs and HDL cholesterol (HDLc) concentrations. Thus, interactions between genes and clinical phenotypes may contribute to this unexplained heritability. We have applied a weighted genetic risk score (GRS) for both plasma TGs and HDLc in two large cohorts at the extremes of BMI. Both BMI and GRS were strongly associated with these lipid traits. A significant interaction between obese/lean status and GRS was noted for each of TG (P(Interaction) = 2.87 × 10(-4)) and HDLc (P(Interaction) = 1.05 × 10(-3)). These interactions were largely driven by SNPs tagging APOA5, glucokinase receptor (GCKR), and LPL for TG, and cholesteryl ester transfer protein (CETP), GalNAc-transferase (GALNT2), endothelial lipase (LIPG), and phospholipid transfer protein (PLTP) for HDLc. In contrast, the GRSLDL cholesterol × adiposity interaction was not significant. Sexual dimorphism was evident for the GRSHDL on HDLc in obese (P(Interaction) = 0.016) but not lean subjects. SNP by BMI interactions may provide biological insight into specific genetic associations and missing heritability.
Keywords: genetic risk score; lipoproteins; obesity; single nucleotide polymorphism; statistical interaction.
Copyright © 2014 by the American Society for Biochemistry and Molecular Biology, Inc.
Figures
References
-
- Johansen C. T., Wang J., Lanktree M. B., McIntyre A. D., Ban M. R., Martins R. A., Kennedy B. A., Hassell R. G., Visser M. E., Schwartz S. M., et al. 2011. An increased burden of common and rare lipid-associated risk alleles contributes to the phenotypic spectrum of hypertriglyceridemia. Arterioscler. Thromb. Vasc. Biol. 31: 1916–1926. - PMC - PubMed
-
- Namboodiri K. K., Kaplan E. B., Heuch I., Elston R. C., Green P. P., Rao D. C., Laskarzewski P., Glueck C. J., Rifkind B. M. 1985. The Collaborative Lipid Research Clinics Family Study: biological and cultural determinants of familial resemblance for plasma lipids and lipoproteins. Genet. Epidemiol. 2: 227–254. - PubMed
-
- Yu Y., Wyszynski D. F., Waterworth D. M., Wilton S. D., Barter P. J., Kesaniemi Y. A., Mahley R. W., McPherson R., Waeber G., Bersot T. P., et al. 2005. Multiple QTLs influencing triglyceride and HDL and total cholesterol levels identified in families with atherogenic dyslipidemia. J. Lipid Res. 46: 2202–2213. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

