Characterization of an alphamesonivirus 3C-like protease defines a special group of nidovirus main proteases
- PMID: 25231310
- PMCID: PMC4248970
- DOI: 10.1128/JVI.02040-14
Characterization of an alphamesonivirus 3C-like protease defines a special group of nidovirus main proteases
Abstract
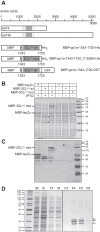
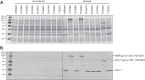
Cavally virus (CavV) and related viruses in the family Mesoniviridae diverged profoundly from other nidovirus lineages but largely retained the characteristic set of replicative enzymes conserved in the Coronaviridae and Roniviridae. The expression of these enzymes in virus-infected cells requires the extensive proteolytic processing of two large replicase polyproteins, pp1a and pp1ab, by the viral 3C-like protease (3CL(pro)). Here, we show that CavV 3CL(pro) autoproteolytic cleavage occurs at two N-terminal (N1 and N2) and one C-terminal (C1) processing site(s). The mature form of 3CL(pro) was revealed to be a 314-residue protein produced by cleavage at FKNK1386|SAAS (N2) and YYNQ1700|SATI (C1). Site-directed mutagenesis data suggest that the mesonivirus 3CL(pro) employs a catalytic Cys-His dyad comprised of CavV pp1a/pp1ab residues Cys-1539 and His-1434. The study further suggests that mesonivirus 3CL(pro) substrate specificities differ from those of related nidovirus proteases. The presence of Gln (or Glu) at the P1 position was not required for cleavage, although residues that control Gln/Glu specificity in related viral proteases are retained in the CavV 3CL(pro) sequence. Asn at the P2 position was identified as a key determinant for mesonivirus 3CL(pro) substrate specificity. Other positions, including P4 and P1', each are occupied by structurally related amino acids, indicating a supportive role in substrate binding. Together, the data identify a new subgroup of nidovirus main proteases and support previous conclusions on phylogenetic relationships between the main nidovirus lineages.
Importance: Mesoniviruses have been suggested to provide an evolutionary link between nidovirus lineages with small (13 to 16 kb) and large (26 to 32 kb) RNA genome sizes, and it has been proposed that a specific set of enzymes, including a proofreading exoribonuclease and other replicase gene-encoded proteins, play a key role in the major genome expansion leading to the currently known lineages of large nidoviruses. Despite their smaller genome size (20 kb), mesoniviruses retained most of the replicative domains conserved in large nidoviruses; thus, they are considered interesting models for studying possible key events in the evolution of RNA genomes of exceptional size and complexity. Our study provides the first characterization of a mesonivirus replicase gene-encoded nonstructural protein. The data confirm and extend previous phylogenetic studies of mesoniviruses and related viruses and pave the way for studies into the formation of the mesonivirus replication complex and functional and structural studies of its functional subunits.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures




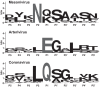
Similar articles
-
Characterization of Self-Processing Activities and Substrate Specificities of Porcine Torovirus 3C-Like Protease.J Virol. 2020 Sep 29;94(20):e01282-20. doi: 10.1128/JVI.01282-20. Print 2020 Sep 29. J Virol. 2020. PMID: 32727876 Free PMC article.
-
Proteolytic processing of mesonivirus replicase polyproteins by the viral 3C-like protease.J Gen Virol. 2016 Jun;97(6):1439-1445. doi: 10.1099/jgv.0.000458. Epub 2016 Mar 15. J Gen Virol. 2016. PMID: 26977900
-
The 3C-like proteinase of an invertebrate nidovirus links coronavirus and potyvirus homologs.J Virol. 2003 Jan;77(2):1415-26. doi: 10.1128/jvi.77.2.1415-1426.2003. J Virol. 2003. PMID: 12502857 Free PMC article.
-
What we know but do not understand about nidovirus helicases.Virus Res. 2015 Apr 16;202:12-32. doi: 10.1016/j.virusres.2014.12.001. Epub 2014 Dec 8. Virus Res. 2015. PMID: 25497126 Free PMC article. Review.
-
Nidovirales: evolving the largest RNA virus genome.Virus Res. 2006 Apr;117(1):17-37. doi: 10.1016/j.virusres.2006.01.017. Epub 2006 Feb 28. Virus Res. 2006. PMID: 16503362 Free PMC article. Review.
Cited by
-
Antiviral activity of K22 against members of the order Nidovirales.Virus Res. 2018 Feb 15;246:28-34. doi: 10.1016/j.virusres.2018.01.002. Epub 2018 Jan 11. Virus Res. 2018. PMID: 29337162 Free PMC article.
-
Isolation and characterization of a novel mesonivirus from Culex mosquitoes in China.Virus Res. 2017 Aug 15;240:130-139. doi: 10.1016/j.virusres.2017.08.001. Epub 2017 Aug 18. Virus Res. 2017. PMID: 28823942 Free PMC article.
-
A planarian nidovirus expands the limits of RNA genome size.PLoS Pathog. 2018 Nov 1;14(11):e1007314. doi: 10.1371/journal.ppat.1007314. eCollection 2018 Nov. PLoS Pathog. 2018. PMID: 30383829 Free PMC article.
-
Characterization of Self-Processing Activities and Substrate Specificities of Porcine Torovirus 3C-Like Protease.J Virol. 2020 Sep 29;94(20):e01282-20. doi: 10.1128/JVI.01282-20. Print 2020 Sep 29. J Virol. 2020. PMID: 32727876 Free PMC article.
-
Recent Progress in Torovirus Molecular Biology.Viruses. 2021 Mar 8;13(3):435. doi: 10.3390/v13030435. Viruses. 2021. PMID: 33800523 Free PMC article. Review.
References
-
- de Groot RJ, Cowley JA, Enjuanes L, Faaberg KS, Perlman S, Rottier PJM, Snijder EJ, Ziebuhr J, Gorbalenya AE. 2012. Order Nidovirales, p 785–795 In King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ. (ed), Virus taxonomy. Elsevier, Amsterdam, the Netherlands.
-
- Cowley JA, Dimmock CM, Spann KM, Walker PJ. 2000. Gill-associated virus of Penaeus monodon prawns: an invertebrate virus with ORF1a and ORF1b genes related to arteri- and coronaviruses. J. Gen. Virol. 81:1473–1484. - PubMed
-
- de Groot RJ, Baker SC, Baric R, Enjuanes L, Gorbalenya AE, Holmes KV, Perlman S, Poon L, Rottier PJM, Talbot PJ, Woo PCY, Ziebuhr J. 2012. Family Coronaviridae, p 806–828 In King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ. (ed), Virus taxonomy. Elsevier, Amsterdam, the Netherlands.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

