Fine-mapping nicotine resistance loci in Drosophila using a multiparent advanced generation inter-cross population
- PMID: 25236448
- PMCID: PMC4174953
- DOI: 10.1534/genetics.114.162107
Fine-mapping nicotine resistance loci in Drosophila using a multiparent advanced generation inter-cross population
Abstract
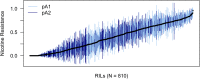
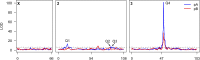
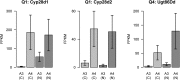
Animals in nature are frequently challenged by toxic compounds, from those that occur naturally in plants as a defense against herbivory, to pesticides used to protect crops. On exposure to such xenobiotic substances, animals mount a transcriptional response, generating detoxification enzymes and transporters that metabolize and remove the toxin. Genetic variation in this response can lead to variation in the susceptibility of different genotypes to the toxic effects of a given xenobiotic. Here we use Drosophila melanogaster to dissect the genetic basis of larval resistance to nicotine, a common plant defense chemical and widely used addictive drug in humans. We identified quantitative trait loci (QTL) for the trait using the DSPR (Drosophila Synthetic Population Resource), a panel of multiparental advanced intercross lines. Mapped QTL collectively explain 68.4% of the broad-sense heritability for nicotine resistance. The two largest-effect loci-contributing 50.3 and 8.5% to the genetic variation-map to short regions encompassing members of classic detoxification gene families. The largest QTL resides over a cluster of ten UDP-glucuronosyltransferase (UGT) genes, while the next largest QTL harbors a pair of cytochrome P450 genes. Using RNAseq we measured gene expression in a pair of DSPR founders predicted to harbor different alleles at both QTL and showed that Ugt86Dd, Cyp28d1, and Cyp28d2 had significantly higher expression in the founder carrying the allele conferring greater resistance. These genes are very strong candidates to harbor causative, regulatory polymorphisms that explain a large fraction of the genetic variation in larval nicotine resistance in the DSPR.
Keywords: DSPR; Drosophila melanogaster; MPP; Multiparent Advanced Generation Inter-Cross (MAGIC); Multiparental populations; QTL mapping; RIL; RNAseq; complex traits; nicotine; quantitative genetics.
Copyright © 2014 by the Genetics Society of America.
Figures

References
-
- Beavis, W. D., 1994 The power and deceit of QTL experiments: lessons from comparative QTL studies. Proceedings of the 49th Annual Corn and Sorghum Industry Research Conference. American Seed Trade Association, Washington, DC, pp. 250–266.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

