Signal peptide peptidase functions in ERAD to cleave the unfolded protein response regulator XBP1u
- PMID: 25239945
- PMCID: PMC4283407
- DOI: 10.15252/embj.201488208
Signal peptide peptidase functions in ERAD to cleave the unfolded protein response regulator XBP1u
Abstract
Signal peptide peptidase (SPP) catalyzes intramembrane proteolysis of signal peptides at the endoplasmic reticulum (ER), but has also been suggested to play a role in ER-associated degradation (ERAD). Here, we show that SPP forms a complex with the ERAD factor Derlin1 and the E3 ubiquitin ligase TRC8 to cleave the unfolded protein response (UPR) regulator XBP1u. Cleavage occurs within a so far unrecognized type II transmembrane domain, which renders XBP1u as an SPP substrate through specific sequence features. Additionally, Derlin1 acts in the complex as a substrate receptor by recognizing the luminal tail of XBP1u. Remarkably, this interaction of Derlin1 with XBP1u obviates the need for ectodomain shedding prior to SPP cleavage, commonly required for intramembrane cuts. Furthermore, we show that XBP1u inhibits the UPR transcription factor XBP1s by targeting it toward proteasomal degradation. Thus, we identify an ERAD complex that controls the abundance of XBP1u and thereby tunes signaling through the UPR.
Keywords: E3 ubiquitin‐protein ligase; GxGD intramembrane protease; protein homeostasis; rhomboid pseudoprotease.
© 2014 The Authors.
Figures

Immunoprecipitation (IP) of CHAPS-solubilized HA-tagged SPP (SPP-HA) and Western blot (WB) analysis with indicated antibodies. Filled triangle, monomer of SPP-HA and endogenous (end.) SPP; open triangle, SDS-stable SPP dimer; asterisks, cross-reacting immunoglobulin chains.
IP/WB analysis of endogenous SPP as in (A).
IP/WB analysis of endogenous Derlin1 as in (A).
BN-PAGE and WB analysis of immunoisolated digitonin-solubilized SPP-HA resolves a 200-kDa (SPP-C200) from a 500-kDa (SPP-C500) complex.
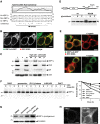
Kyte-Doolittle hydrophobicity plot (window size of 5) of human (H. sapiens, Hs) XBP1u and comparison to its mouse (M. musculus, Mm) and zebrafish (D. rerio, Dr) orthologues reveals a conserved low-scoring TM domain.
XBP1u N-terminally tagged with GFP co-localized with the ER marker RFP-KDEL. Scale bar, 5 μm.
Subcellular fractionation, extraction of the microsomal membrane fraction (m) with high salt and sodium carbonate reveals that XBP1u-N-FLAG is an integral membrane protein. t, total cell extract; c, cytosol; sn, supernatant, p, pellet.
XBP1u is a type II membrane protein as demonstrated by the sensitivity of the glycosylated R232N mutant (open triangle) to EndoH (H) and PNGaseF (P).
FPP assay shows that GFP fused to the C-terminus of XBP1u is protected, whereas an N-terminal fusion is accessible for trypsin treatment. The luminal ER marker RFP-KDEL and CD3δ with mCherry fused to its cytosolic C-terminus were used as positive and negative controls, respectively. Of note, low-level nuclear GFP signal was not present when SPP-catalyzed cleavage of XBP1u was blocked (see Fig 4B and Supplementary Fig S3), indicating that it is a consequence of SPP-triggered release. Scale bar, 5 μm.
Cycloheximide (CHX) chase in presence of 5 μM epoxomicin, 50 μM (Z-LL)2-ketone, 5 μM L-685,458, and 50 μM DAPT, respectively. Western blot quantification is shown (means ± SEM, n = 3). DMSO, vehicle control; asterisks, N-terminal XBP1u fragment.
Treatment of HEK293T cells with 5 μM epoxomicin, 50 μM (Z-LL)2-ketone, and 5 μM L-685,458 for 16 h, respectively, and analysis of endogenous XBP1u in isolated microsomal membrane fractions. The unrelated structural ER protein CLIMP63 was used as a loading control.
Immunofluorescence analysis of HEK293T cells shows that endogenous XBP1u accumulates in cells treated with SPP inhibitor L-685,458 (5 μM). Scale bar, 5 μm.

The primary structure of the XBP1u TM domain determines degradation kinetics as assessed by cycloheximide (CHX) chase and Western blotting (means ± SEM, n = 3).
CD spectra of XBP1u TM domain peptides tagged at both termini with KKK sequences (see Supplementary Materials and Methods) and were obtained in 40% trifluoroethanol in aqueous buffer which mimics the water-filled lumen of an intramembrane protease while preventing unspecific peptide aggregation (means ± SD, n = 3). Inset: secondary structure contents as calculated from the CD spectra.
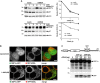
Doxycycline-induced expression of p97-specific knockdown construct (shRNA) stabilized FLAG-tagged CD3δ but had no influence on XBP1u-N-FLAG as assessed by cycloheximide (CHX) chase. Right panel, Western blot quantification (means ± SEM, n = 3).
Fluorescence microscopy of XBP1u-GFP- and RFP-KDEL-expressing cells that had been treated with 0.5 μM epoxomicin and 5 μM L-685,458, respectively. Scale bars, 5 μm.
Co-expression of XBP1u-N-FLAG with SPP WT in presence of 5 μM epoxomicin increased levels of N-terminal fragment (open triangle), whereas the catalytically inactive SPPD265A mutant shows no activity.
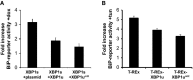
Doxycycline (dox)-induced XBP1s expression in HEK293 T-REx cells induces BiP-reporter activity. Co-expression of XBP1u and the XBP1umt2 reduces this activation. Quantification of relative induction of luciferase activity is shown (means ± SEM, n = 3).
Doxycycline-induced expression of XBP1u and XBP1umt2 in HEK293 T-REx cells reduces tunicamycin (tun)-induced activation of the endogenous UPR compared to the parenteral T-REx cells as assessed by a BiP-reporter luciferase assay (means ± SEM, n = 3).

Co-expression of SPPD265A stabilized XBP1u, whereas YOD1C160S and SPPL3DAKK had no effect. Right panel, quantification of the relative amount of 35S-pulse-labeled XBP1u-N-FLAG (means ± SEM, n = 3). Asterisks, unidentified co-immunoprecipitated protein.
HA-tagged SPP (SPP-HA) and the catalytic mutant (SPP-DA-HA) co-immunoprecipitate with XBP1u but not with RAMP4.
Substrate-trapping SPP-DA-HA was co-expressed with XBP1u (left panel) or OP91* (right panel) and analyzed by BN-PAGE as shown (Fig 1D).

Cells transfected with XBP1u-N-FLAG were co-transfected either with empty vector (mock), SPPD265A, TRC8ΔR or Derlin1G180V as indicated and subjected to 35S-pulse-label chase analysis (means ± SEM, n = 3). Asterisks, unidentified co-immunoprecipitated protein.
Pulse-chase analysis of XBP1uΔ turnover as in (A).
GFP-tagged Derlin1G180V co-immunoprecipitates with FLAG-tagged XBP1u but not with XBP1uΔ.
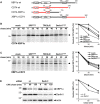
Outline of domain fusions between CD74 (indicated in red) and XBP1u.
CD74-XBP1u chimera was co-transfected either with empty vector (mock), SPPD265A, TRC8ΔR or Derlin1G180V as indicated and subjected to 35S-pulse-label chase analysis (means ± SEM, n = 3).
Pulse-chase analysis of XBP1u-CD74 turnover as in (B).
Knockdown of Derlin1 by siRNA reduces XBP1u degradation (means ± SEM, n = 3). CHX, cycloheximide chase; nt, non-targeting control siRNA.

References
-
- Bagola K, Mehnert M, Jarosch E, Sommer T. Protein dislocation from the ER. Biochim Biophys Acta. 2011;1808:925–936. - PubMed
-
- Beisner DR, Langerak P, Parker AE, Dahlberg C, Otero FJ, Sutton SE, Poirot L, Barnes W, Young MA, Niessen S, Wiltshire T, Bodendorf U, Martoglio B, Cravatt B, Cooke MP. The intramembrane protease Sppl2a is required for B cell and DC development and survival via cleavage of the invariant chain. J Exp Med. 2013;210:23–30. - PMC - PubMed
-
- Bergmann H, Yabas M, Short A, Miosge L, Barthel N, Teh CE, Roots CM, Bull KR, Jeelall Y, Horikawa K, Whittle B, Balakishnan B, Sjollema G, Bertram EM, Mackay F, Rimmer AJ, Cornall RJ, Field MA, Andrews TD, Goodnow CC, et al. B cell survival, surface BCR and BAFFR expression, CD74 metabolism, and CD8- dendritic cells require the intramembrane endopeptidase SPPL2A. J Exp Med. 2013;210:31–40. - PMC - PubMed
-
- Brauweiler A, Lorick KL, Lee JP, Tsai YC, Chan D, Weissman AM, Drabkin HA, Gemmill RM. RING-dependent tumor suppression and G2/M arrest induced by the TRC8 hereditary kidney cancer gene. Oncogene. 2007;26:2263–2271. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

