Interactions Among Plant Transcription Factors Regulating Expression of Stress-responsive Genes
- PMID: 25249757
- PMCID: PMC4167486
- DOI: 10.4137/BBI.S16313
Interactions Among Plant Transcription Factors Regulating Expression of Stress-responsive Genes
Abstract
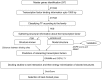
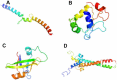
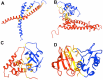
Plants are simultaneously subjected to a variety of stress conditions in the field and are known to combat the hostile conditions by up/down-regulating number of genes. There exists a significant level of cross-talk between different stress responses in plants. In this study, we predict the interacting pairs of transcription factors that regulate the multiple abiotic stress-responsive genes in the plant Arabidopsis thaliana. We identified the interacting pair(s) of transcription factors (TFs) based on the spatial proximity of their binding sites. We also examined the interactions between the predicted pairs of TFs using molecular docking. Subsequent to docking, the best interaction pose was selected using our scoring scheme DockScore, which ranks the docked solutions based on several interface parameters and aims to find optimal interactions between proteins. We analyzed the selected docked pose for the interface residues and their conservation.
Keywords: Arabidopsis thaliana; docking; protein–protein interactions; stress conditions; transcription factors.
Figures
Similar articles
-
Structural interface parameters are discriminatory in recognising near-native poses of protein-protein interactions.PLoS One. 2014 Feb 3;9(2):e80255. doi: 10.1371/journal.pone.0080255. eCollection 2014. PLoS One. 2014. PMID: 24498255 Free PMC article.
-
DOCKSCORE: a webserver for ranking protein-protein docked poses.BMC Bioinformatics. 2015 Apr 24;16(1):127. doi: 10.1186/s12859-015-0572-6. BMC Bioinformatics. 2015. PMID: 25902779 Free PMC article.
-
Identification of transcription factor's targets using tissue-specific transcriptomic data in Arabidopsis thaliana.BMC Syst Biol. 2010 Sep 13;4 Suppl 2(Suppl 2):S2. doi: 10.1186/1752-0509-4-S2-S2. BMC Syst Biol. 2010. PMID: 20840729 Free PMC article.
-
From experiment-driven database analyses to database-driven experiments in Arabidopsis thaliana transcription factor research.Plant Sci. 2017 Sep;262:141-147. doi: 10.1016/j.plantsci.2017.06.011. Epub 2017 Jun 27. Plant Sci. 2017. PMID: 28716409 Review.
-
Revisiting the Role of Plant Transcription Factors in the Battle against Abiotic Stress.Int J Mol Sci. 2018 May 31;19(6):1634. doi: 10.3390/ijms19061634. Int J Mol Sci. 2018. PMID: 29857524 Free PMC article. Review.
Cited by
-
Stress-Mediated cis-Element Transcription Factor Interactions Interconnecting Primary and Specialized Metabolism in planta.Front Plant Sci. 2016 Nov 25;7:1725. doi: 10.3389/fpls.2016.01725. eCollection 2016. Front Plant Sci. 2016. PMID: 27933071 Free PMC article. Review.
-
Identification and expression analysis of the DREB transcription factor family in pineapple (Ananas comosus (L.) Merr.).PeerJ. 2020 Apr 28;8:e9006. doi: 10.7717/peerj.9006. eCollection 2020. PeerJ. 2020. PMID: 32377449 Free PMC article.
-
Regulation of a single inositol 1-phosphate synthase homeologue by HSFA6B contributes to fibre yield maintenance under drought conditions in upland cotton.Plant Biotechnol J. 2024 Oct;22(10):2756-2772. doi: 10.1111/pbi.14402. Epub 2024 Jun 21. Plant Biotechnol J. 2024. PMID: 39031479 Free PMC article.
-
Network and biosignature analysis for the integration of transcriptomic and metabolomic data to characterize leaf senescence process in sunflower.BMC Bioinformatics. 2016 Jun 6;17 Suppl 5(Suppl 5):174. doi: 10.1186/s12859-016-1045-2. BMC Bioinformatics. 2016. PMID: 27295368 Free PMC article.
-
Physiological and Transcriptomic Responses of Chinese Cabbage (Brassica rapa L. ssp. Pekinensis) to Salt Stress.Int J Mol Sci. 2017 Sep 12;18(9):1953. doi: 10.3390/ijms18091953. Int J Mol Sci. 2017. PMID: 28895882 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

