Alternatively spliced transcripts and novel pseudogenes of the Plasmodium falciparum resistance-associated locus pfcrt detected in East African malaria patients
- PMID: 25253286
- PMCID: PMC4267505
- DOI: 10.1093/jac/dku358
Alternatively spliced transcripts and novel pseudogenes of the Plasmodium falciparum resistance-associated locus pfcrt detected in East African malaria patients
Abstract
Objectives: Polymorphisms in the lysosomal transporter encoded by the pfcrt gene directly impact on Plasmodium falciparum susceptibility to aminoquinolines. The Lys76Thr mutation is the critical change conferring chloroquine resistance in vitro and in vivo, but always occurs with additional non-synonymous changes in the pfcrt coding sequence. We sought to better describe pfcrt polymorphisms distal to codon 76.
Methods: Small-volume samples (≤ 500 μL) of parasite-infected blood collected directly from malaria patients presenting for treatment in Sudan and Tanzania were immediately preserved for RNA extraction. The pfcrt locus was amplified from cDNA preparations by nested PCR and sequenced directly to derive full-length mRNA sequences.
Results: In one of two sites in Sudan, two patients were found with an unorthodox spliced form of pfcrt mRNA in which two exons were skipped, but it was not possible to test for the presence of the putative protein products of these aberrant transcripts. Genomic DNA sequencing from dried blood spots collected in parallel confirmed the presence of spliced pfcrt pseudogenes in a minority of parasite isolates. Full-length cDNA from conventionally spliced mRNA molecules in all study sites demonstrated the existence of a variety of pfcrt haplotypes in East Africa, and thus provides evidence of intragenic recombination.
Conclusions: The presence of pseudogenes, although unlikely to have any direct public health impact, may confound results obtained from simple genotyping methods that consider only codon 76 and the adjacent residues of pfcrt.
Keywords: alternative splicing; chloroquine resistance transporters; malaria.
© The Author 2014. Published by Oxford University Press on behalf of the British Society for Antimicrobial Chemotherapy.
Figures



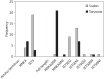

References
-
- White NJ, Nosten F, Looareesuwan S, et al. Averting a malaria disaster. Lancet. 1999;353:1965–7. - PubMed
-
- Breman JG. Resistance to artemisinin-based combination therapy. Lancet Infect Dis. 2012;12:820–2. - PubMed
-
- Noedl H, Se Y, Schaecher K, et al. Evidence of artemisinin-resistant malaria in western Cambodia. New Engl J Med. 2008;359:2619–20. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

