Identification of a DNA methylation signature to predict disease-free survival in locally advanced rectal cancer
- PMID: 25261372
- PMCID: PMC4226671
- DOI: 10.18632/oncotarget.2347
Identification of a DNA methylation signature to predict disease-free survival in locally advanced rectal cancer
Abstract
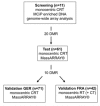
In locally advanced rectal cancer a preoperative predictive biomarker is necessary to adjust treatment specifically for those patients expected to suffer relapse. We applied whole genome methylation CpG island array analyses to an initial set of patients (n=11) to identify differentially methylated regions (DMRs) that separate a good from a bad prognosis group. Using a quantitative high-resolution approach, candidate DMRs were first validated in a set of 61 patients (test set) and then confirmed DMRs were further validated in additional independent patient cohorts (n=71, n=42). We identified twenty highly discriminative DMRs and validated them in the test set using the MassARRAY technique. Ten DMRs could be confirmed which allowed separation into prognosis groups (p=0.0207, HR=4.09). The classifier was validated in two additional cohorts (n=71, p=0.0345, HR=3.57 and n=42, p=0.0113, HR=3.78). Interestingly, six of the ten DMRs represented regions close to the transcriptional start sites of genes which are also marked by the Polycomb Repressor Complex component EZH2. In conclusion we present a classifier comprising 10 DMRs which predicts patient prognosis with a high degree of accuracy. These data may now help to discriminate between patients that may respond better to standard treatments from those that may require alternative modalities.
Conflict of interest statement
No potential conflicts of interest were disclosed for any oft he authors.
Figures


Similar articles
-
Genome-wide analysis of CpG island methylation in bladder cancer identified TBX2, TBX3, GATA2, and ZIC4 as pTa-specific prognostic markers.Eur Urol. 2012 Jun;61(6):1245-56. doi: 10.1016/j.eururo.2012.01.011. Epub 2012 Jan 18. Eur Urol. 2012. PMID: 22284968
-
Circulating serum microRNA-345 correlates with unfavorable pathological response to preoperative chemoradiotherapy in locally advanced rectal cancer.Oncotarget. 2016 Sep 27;7(39):64233-64243. doi: 10.18632/oncotarget.11649. Oncotarget. 2016. PMID: 27572313 Free PMC article.
-
Integrated analysis of DNA methylation profiling and gene expression profiling identifies novel markers in lung cancer in Xuanwei, China.PLoS One. 2018 Oct 4;13(10):e0203155. doi: 10.1371/journal.pone.0203155. eCollection 2018. PLoS One. 2018. PMID: 30286088 Free PMC article.
-
Gene expression classification of colon cancer into molecular subtypes: characterization, validation, and prognostic value.PLoS Med. 2013;10(5):e1001453. doi: 10.1371/journal.pmed.1001453. Epub 2013 May 21. PLoS Med. 2013. PMID: 23700391 Free PMC article.
-
Analyzing the cancer methylome through targeted bisulfite sequencing.Cancer Lett. 2013 Nov 1;340(2):171-8. doi: 10.1016/j.canlet.2012.10.040. Epub 2012 Nov 28. Cancer Lett. 2013. PMID: 23200671 Free PMC article. Review.
Cited by
-
Development of a 15-gene signature for predicting prognosis in advanced colorectal cancer.Bioengineered. 2020 Dec;11(1):165-174. doi: 10.1080/21655979.2020.1718459. Bioengineered. 2020. PMID: 32036725 Free PMC article.
-
Epigenetic Biomarkers in Colorectal Cancer Patients Receiving Adjuvant or Neoadjuvant Therapy: A Systematic Review of Epidemiological Studies.Int J Mol Sci. 2019 Aug 6;20(15):3842. doi: 10.3390/ijms20153842. Int J Mol Sci. 2019. PMID: 31390840 Free PMC article.
-
Cell-free plasma hypermethylated CASZ1, CDH13 and ING2 are promising biomarkers of esophageal cancer.J Biomed Res. 2018 Nov 20;32(5):424-433. doi: 10.7555/JBR.32.20170065. J Biomed Res. 2018. PMID: 30355852 Free PMC article.
-
Prognostic Value of MicroRNAs in Preoperative Treated Rectal Cancer.Int J Mol Sci. 2016 Apr 15;17(4):568. doi: 10.3390/ijms17040568. Int J Mol Sci. 2016. PMID: 27092493 Free PMC article.
-
Integrating Tumor and Organoid DNA Methylation Profiles Reveals Robust Predictors of Chemotherapy Response in Rectal Cancer.medRxiv [Preprint]. 2025 Mar 4:2025.02.28.25322951. doi: 10.1101/2025.02.28.25322951. medRxiv. 2025. PMID: 40093220 Free PMC article. Preprint.
References
-
- Engstrom PF, Arnoletti JP, Benson AB, 3rd, Chen YJ, Choti MA, Cooper HS, Covey A, Dilawari RA, Early DS, Enzinger PC, Fakih MG, Fleshman J, Jr, Fuchs C, Grem JL, Kiel K, Knol JA, et al. NCCN Clinical Practice Guidelines in Oncology: rectal cancer. J Natl Compr Canc Netw. 2009;7:838–881. - PubMed
-
- Schmiegel W, Reinacher-Schick A, Arnold D, Graeven U, Heinemann V, Porschen R, Riemann J, Rodel C, Sauer R, Wieser M, Schmitt W, Schmoll HJ, Seufferlein T, Kopp I, Pox C. [Update S3-guideline “colorectal cancer” 2008] Zeitschrift fur Gastroenterologie. 2008;46:799–840. - PubMed
-
- Gerard JP, Conroy T, Bonnetain F, Bouche O, Chapet O, Closon-Dejardin MT, Untereiner M, Leduc B, Francois E, Maurel J, Seitz JF, Buecher B, Mackiewicz R, Ducreux M, Bedenne L. Preoperative radiotherapy with or without concurrent fluorouracil and leucovorin in T3–4 rectal cancers: results of FFCD 9203. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2006;24:4620–4625. - PubMed
-
- Sauer R, Becker H, Hohenberger W, Rodel C, Wittekind C, Fietkau R, Martus P, Tschmelitsch J, Hager E, Hess CF, Karstens JH, Liersch T, Schmidberger H, Raab R. Preoperative versus postoperative chemoradiotherapy for rectal cancer. The New England journal of medicine. 2004;351:1731–1740. - PubMed
-
- Sauer R, Liersch T, Merkel S, Fietkau R, Hohenberger W, Hess C, Becker H, Raab HR, Villanueva MT, Witzigmann H, Wittekind C, Beissbarth T, Rodel C. Preoperative Versus Postoperative Chemoradiotherapy for Locally Advanced Rectal Cancer: Results of the German CAO/ARO/AIO-94 Randomized Phase III Trial After a Median Follow-Up of 11 Years. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2012;30:1926–1933. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

