CRISPR-Cas9 knockin mice for genome editing and cancer modeling
- PMID: 25263330
- PMCID: PMC4265475
- DOI: 10.1016/j.cell.2014.09.014
CRISPR-Cas9 knockin mice for genome editing and cancer modeling
Abstract
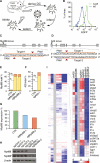
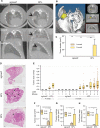
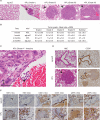
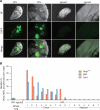
CRISPR-Cas9 is a versatile genome editing technology for studying the functions of genetic elements. To broadly enable the application of Cas9 in vivo, we established a Cre-dependent Cas9 knockin mouse. We demonstrated in vivo as well as ex vivo genome editing using adeno-associated virus (AAV)-, lentivirus-, or particle-mediated delivery of guide RNA in neurons, immune cells, and endothelial cells. Using these mice, we simultaneously modeled the dynamics of KRAS, p53, and LKB1, the top three significantly mutated genes in lung adenocarcinoma. Delivery of a single AAV vector in the lung generated loss-of-function mutations in p53 and Lkb1, as well as homology-directed repair-mediated Kras(G12D) mutations, leading to macroscopic tumors of adenocarcinoma pathology. Together, these results suggest that Cas9 mice empower a wide range of biological and disease modeling applications.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures



References
-
- Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315:1709–1712. - PubMed
Publication types
MeSH terms
Associated data
Grants and funding
- P30 DK043351/DK/NIDDK NIH HHS/United States
- HHMI/Howard Hughes Medical Institute/United States
- U54CA151884/CA/NCI NIH HHS/United States
- P30-CA14051/CA/NCI NIH HHS/United States
- P50 HG006193/HG/NHGRI NIH HHS/United States
- R01 DK097768/DK/NIDDK NIH HHS/United States
- R37 EB000244/EB/NIBIB NIH HHS/United States
- EB000244/EB/NIBIB NIH HHS/United States
- R01-NS 07312401/NS/NINDS NIH HHS/United States
- R01-CA133404/CA/NCI NIH HHS/United States
- U54 CA151884/CA/NCI NIH HHS/United States
- P30 CA014051/CA/NCI NIH HHS/United States
- DP1-MH100706/DP/NCCDPHP CDC HHS/United States
- DK43351/DK/NIDDK NIH HHS/United States
- R01 CA133404/CA/NCI NIH HHS/United States
- DP1 MH100706/MH/NIMH NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

