Histone deacetylase inhibition rescues structural and functional brain deficits in a mouse model of Kabuki syndrome
- PMID: 25273096
- PMCID: PMC4406328
- DOI: 10.1126/scitranslmed.3009278
Histone deacetylase inhibition rescues structural and functional brain deficits in a mouse model of Kabuki syndrome
Abstract
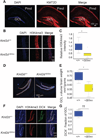
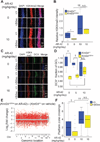
Kabuki syndrome is caused by haploinsufficiency for either of two genes that promote the opening of chromatin. If an imbalance between open and closed chromatin is central to the pathogenesis of Kabuki syndrome, agents that promote chromatin opening might have therapeutic potential. We have characterized a mouse model of Kabuki syndrome with a heterozygous deletion in the gene encoding the lysine-specific methyltransferase 2D (Kmt2d), leading to impairment of methyltransferase function. In vitro reporter alleles demonstrated a reduction in histone 4 acetylation and histone 3 lysine 4 trimethylation (H3K4me3) activity in mouse embryonic fibroblasts from Kmt2d(+/βGeo) mice. These activities were normalized in response to AR-42, a histone deacetylase inhibitor. In vivo, deficiency of H3K4me3 in the dentate gyrus granule cell layer of Kmt2d(+/βGeo) mice correlated with reduced neurogenesis and hippocampal memory defects. These abnormalities improved upon postnatal treatment with AR-42. Our work suggests that a reversible deficiency in postnatal neurogenesis underlies intellectual disability in Kabuki syndrome.
Copyright © 2014, American Association for the Advancement of Science.
Figures

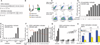
References
-
- Ng SB, Bigham AW, Buckingham KJ, Hannibal MC, McMillin MJ, Gildersleeve HI, Beck AE, Tabor HK, Cooper GM, Mefford HC, Lee C, Turner EH, Smith JD, Rieder MJ, Yoshiura K, Matsumoto N, Ohta T, Niikawa N, Nickerson DA, Bamshad MJ, Shendure J. Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome. Nat. Genet. 2010;42:790–793. - PMC - PubMed
-
- Miyake N, Mizuno S, Okamoto N, Ohashi H, Shiina M, Ogata K, Tsurusaki Y, Nakashima M, Saitsu H, Niikawa N, Matsumoto N. KDM6A point mutations cause Kabuki syndrome. Hum. Mutat. 2013;34:108–110. - PubMed
-
- Altman J. Are new neurons formed in the brains of adult mammals? Science. 1962;135:1127–1128. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

