Visualization and quantification of nascent RAD51 filament formation at single-monomer resolution
- PMID: 25288749
- PMCID: PMC4210327
- DOI: 10.1073/pnas.1307824111
Visualization and quantification of nascent RAD51 filament formation at single-monomer resolution
Abstract
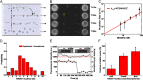
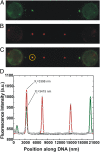
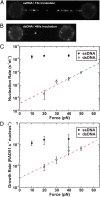
During recombinational repair of double-stranded DNA breaks, RAD51 recombinase assembles as a nucleoprotein filament around single-stranded DNA to form a catalytically proficient structure able to promote homology recognition and strand exchange. Mediators and accessory factors guide the action and control the dynamics of RAD51 filaments. Elucidation of these control mechanisms necessitates development of approaches to quantitatively probe transient aspects of RAD51 filament dynamics. Here, we combine fluorescence microscopy, optical tweezers, and microfluidics to visualize the assembly of RAD51 filaments on bare single-stranded DNA and quantify the process with single-monomer sensitivity. We show that filaments are seeded from RAD51 nuclei that are heterogeneous in size. This heterogeneity appears to arise from the energetic balance between RAD51 self-assembly in solution and the size-dependent interaction time of the nuclei with DNA. We show that nucleation intrinsically is substrate selective, strongly favoring filament formation on bare single-stranded DNA. Furthermore, we devised a single-molecule fluorescence recovery after photobleaching assay to independently observe filament nucleation and growth, permitting direct measurement of their contributions to filament formation. Our findings yield a comprehensive, quantitative understanding of RAD51 filament formation on bare single-stranded DNA that will serve as a basis to elucidate how mediators help RAD51 filament assembly and accessory factors control filament dynamics.
Keywords: BRCA2; RAD51; homologous recombination; optical tweezers; single-molecule fluorescence.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
-
- Hoeijmakers JH. Genome maintenance mechanisms for preventing cancer. Nature. 2001;411(6835):366–374. - PubMed
-
- San Filippo J, Sung P, Klein H. Mechanism of eukaryotic homologous recombination. Annu Rev Biochem. 2008;77:229–257. - PubMed
-
- Sung P, Klein H. Mechanism of homologous recombination: Mediators and helicases take on regulatory functions. Nat Rev Mol Cell Biol. 2006;7(10):739–750. - PubMed
-
- Wyman C, Kanaar R. DNA double-strand break repair: All’s well that ends well. Annu Rev Genet. 2006;40:363–383. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

