Preemptive clinical pharmacogenetics implementation: current programs in five US medical centers
- PMID: 25292429
- PMCID: PMC4607278
- DOI: 10.1146/annurev-pharmtox-010814-124835
Preemptive clinical pharmacogenetics implementation: current programs in five US medical centers
Abstract
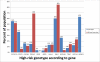
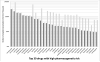
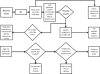
Although the field of pharmacogenetics has existed for decades, practioners have been slow to implement pharmacogenetic testing in clinical care. Numerous publications describe the barriers to clinical implementation of pharmacogenetics. Recently, several freely available resources have been developed to help address these barriers. In this review, we discuss current programs that use preemptive genotyping to optimize the pharmacotherapy of patients. Array-based preemptive testing includes a large number of relevant pharmacogenes that impact multiple high-risk drugs. Using a preemptive approach allows genotyping results to be available prior to any prescribing decision so that genomic variation may be considered as an inherent patient characteristic in the planning of therapy. This review describes the common elements among programs that have implemented preemptive genotyping and highlights key processes for implementation, including clinical decision support.
Keywords: clinical decision support; individualized medicine; personalized medicine; pharmacogenomics; precision medicine; prediction in pharmacology.
Figures

References
-
- Meyer UA. Pharmacogenetics - five decades of therapeutic lessons from genetic diversity. Nat Rev Genet. 2004;5:669–76. - PubMed
-
- Green ED, Guyer MS. Charting a course for genomic medicine from base pairs to bedside. Nature. 2011;470:204–13. - PubMed
-
- Manolio TA, Green ED. Genomics reaches the clinic: from basic discoveries to clinical impact. Cell. 2011;147:14–6. - PubMed
-
- Voora D, Ginsburg GS. A hub for bench-to-bedside pharmacogenomic-based research. Pharmacogenomics. 2011;12:1095–8. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

