The T-box gene family: emerging roles in development, stem cells and cancer
- PMID: 25294936
- PMCID: PMC4197708
- DOI: 10.1242/dev.104471
The T-box gene family: emerging roles in development, stem cells and cancer
Abstract
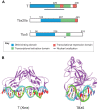
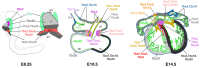
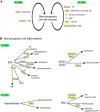
The T-box family of transcription factors exhibits widespread involvement throughout development in all metazoans. T-box proteins are characterized by a DNA-binding motif known as the T-domain that binds DNA in a sequence-specific manner. In humans, mutations in many of the genes within the T-box family result in developmental syndromes, and there is increasing evidence to support a role for these factors in certain cancers. In addition, although early studies focused on the role of T-box factors in early embryogenesis, recent studies in mice have uncovered additional roles in unsuspected places, for example in adult stem cell populations. Here, I provide an overview of the key features of T-box transcription factors and highlight their roles and mechanisms of action during various stages of development and in stem/progenitor cell populations.
Keywords: T-box genes; Tbx; Transcription factors.
© 2014. Published by The Company of Biologists Ltd.
Figures




References
-
- Aanhaanen, W. T. J., Boukens, B. J. D., Sizarov, A., Wakker, V., de Gier-de Vries, C., van Ginneken, A. C., Moorman, A. F. M., Coronel, R. and Christoffels, V. M. (2011). Defective Tbx2-dependent patterning of the atrioventricular canal myocardium causes accessory pathway formation in mice. J. Clin. Invest. 121, 534-544 10.1172/JCI44350 - DOI - PMC - PubMed
-
- Abrahams, A., Parker, M. I. and Prince, S. (2010). The T-box transcription factor Tbx2: its role in development and possible implication in cancer. IUBMB Life 62, 92-102. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

