A customizable software for fast reduction and analysis of large X-ray scattering data sets: applications of the new DPDAK package to small-angle X-ray scattering and grazing-incidence small-angle X-ray scattering
- PMID: 25294982
- PMCID: PMC4180741
- DOI: 10.1107/S1600576714019773
A customizable software for fast reduction and analysis of large X-ray scattering data sets: applications of the new DPDAK package to small-angle X-ray scattering and grazing-incidence small-angle X-ray scattering
Abstract
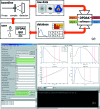
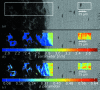
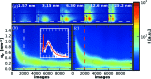
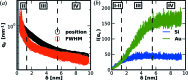
X-ray scattering experiments at synchrotron sources are characterized by large and constantly increasing amounts of data. The great number of files generated during a synchrotron experiment is often a limiting factor in the analysis of the data, since appropriate software is rarely available to perform fast and tailored data processing. Furthermore, it is often necessary to perform online data reduction and analysis during the experiment in order to interactively optimize experimental design. This article presents an open-source software package developed to process large amounts of data from synchrotron scattering experiments. These data reduction processes involve calibration and correction of raw data, one- or two-dimensional integration, as well as fitting and further analysis of the data, including the extraction of certain parameters. The software, DPDAK (directly programmable data analysis kit), is based on a plug-in structure and allows individual extension in accordance with the requirements of the user. The article demonstrates the use of DPDAK for on- and offline analysis of scanning small-angle X-ray scattering (SAXS) data on biological samples and microfluidic systems, as well as for a comprehensive analysis of grazing-incidence SAXS data. In addition to a comparison with existing software packages, the structure of DPDAK and the possibilities and limitations are discussed.
Keywords: computer programs; data analysis; data reduction; grazing-incidence small-angle X-ray scattering; small-angle X-ray scattering.
Figures


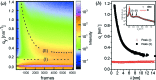

References
-
- Chourou, S. T., Sarje, A., Li, X. S., Chan, E. R. & Hexemer, A. (2013). J. Appl. Cryst. 46, 1781–1795.
-
- Förster, S., Apostol, L. & Bras, W. (2010). J. Appl. Cryst. 43, 639–646.
-
- Fratzl, P., Schreiber, S. & Boyde, A. (1996). Calcif. Tissue Int. 58, 341–346. - PubMed
-
- Fratzl, P., Schreiber, S. & Klaushofer, K. (1996). Connect. Tissue Res. 35, 9–16. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
