The X-ray crystal structure of the euryarchaeal RNA polymerase in an open-clamp configuration
- PMID: 25311937
- PMCID: PMC4657547
- DOI: 10.1038/ncomms6132
The X-ray crystal structure of the euryarchaeal RNA polymerase in an open-clamp configuration
Abstract
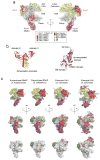
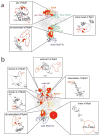
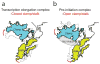
The archaeal transcription apparatus is closely related to the eukaryotic RNA polymerase II (Pol II) system. Archaeal RNA polymerase (RNAP) and Pol II evolved from a common ancestral structure and the euryarchaeal RNAP is the simplest member of the extant archaeal-eukaryotic RNAP family. Here we report the first crystal structure of euryarchaeal RNAP from Thermococcus kodakarensis (Tko). This structure reveals that the clamp domain is able to swing away from the main body of RNAP in the presence of the Rpo4/Rpo7 stalk by coordinated movements of these domains. More detailed structure-function analysis of yeast Pol II and Tko RNAP identifies structural additions to Pol II that correspond to the binding sites of Pol II-specific general transcription factors including TFIIF, TFIIH and Mediator. Such comparisons provide a framework for dissecting interactions between RNAP and these factors during formation of the pre-initiation complex.
Conflict of interest statement
Figures



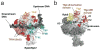

Similar articles
-
Direct binding of TFEα opens DNA binding cleft of RNA polymerase.Nat Commun. 2020 Nov 30;11(1):6123. doi: 10.1038/s41467-020-19998-x. Nat Commun. 2020. PMID: 33257704 Free PMC article.
-
Crystal structure of a pol alpha family DNA polymerase from the hyperthermophilic archaeon Thermococcus sp. 9 degrees N-7.J Mol Biol. 2000 Jun 2;299(2):447-62. doi: 10.1006/jmbi.2000.3728. J Mol Biol. 2000. PMID: 10860752
-
Evolution of complex RNA polymerases: the complete archaeal RNA polymerase structure.PLoS Biol. 2009 May;7(5):e1000102. doi: 10.1371/journal.pbio.1000102. Epub 2009 May 5. PLoS Biol. 2009. PMID: 19419240 Free PMC article.
-
Archaeal RNA polymerase: the influence of the protruding stalk in crystal packing and preliminary biophysical analysis of the Rpo13 subunit.Biochem Soc Trans. 2011 Jan;39(1):25-30. doi: 10.1042/BST0390025. Biochem Soc Trans. 2011. PMID: 21265742 Review.
-
Archaeal RNA polymerase and transcription regulation.Crit Rev Biochem Mol Biol. 2011 Feb;46(1):27-40. doi: 10.3109/10409238.2010.538662. Crit Rev Biochem Mol Biol. 2011. PMID: 21250781 Free PMC article. Review.
Cited by
-
Coupling of Transcription and Translation in Archaea: Cues From the Bacterial World.Front Microbiol. 2021 Apr 29;12:661827. doi: 10.3389/fmicb.2021.661827. eCollection 2021. Front Microbiol. 2021. PMID: 33995325 Free PMC article.
-
Archaeal transcription.Transcription. 2020 Oct;11(5):199-210. doi: 10.1080/21541264.2020.1838865. Epub 2020 Oct 28. Transcription. 2020. PMID: 33112729 Free PMC article. Review.
-
Direct binding of TFEα opens DNA binding cleft of RNA polymerase.Nat Commun. 2020 Nov 30;11(1):6123. doi: 10.1038/s41467-020-19998-x. Nat Commun. 2020. PMID: 33257704 Free PMC article.
-
Widespread formation of alternative 3' UTR isoforms via transcription termination in archaea.Nat Microbiol. 2016 Aug 22;1(10):16143. doi: 10.1038/nmicrobiol.2016.143. Nat Microbiol. 2016. PMID: 27670118
-
The cutting edge of archaeal transcription.Emerg Top Life Sci. 2018 Dec 14;2(4):517-533. doi: 10.1042/ETLS20180014. Emerg Top Life Sci. 2018. PMID: 33525828 Free PMC article.
References
-
- Beattie TR, Bell SD. Molecular machines in archaeal DNA replication. Curr Opin Chem Biol. 2011;15:614–9. - PubMed
-
- Werner F, Grohmann D. Evolution of multisubunit RNA polymerases in the three domains of life. Nat Rev Microbiol. 2011;9:85–98. - PubMed
-
- Fernandez-Tornero C, et al. Crystal structure of the 14-subunit RNA polymerase I. Nature. 2013;502:644–9. - PubMed
-
- Engel C, Sainsbury S, Cheung AC, Kostrewa D, Cramer P. RNA polymerase I structure and transcription regulation. Nature. 2013;502:650–5. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

