Postzygotic single-nucleotide mosaicisms in whole-genome sequences of clinically unremarkable individuals
- PMID: 25312340
- PMCID: PMC4220156
- DOI: 10.1038/cr.2014.131
Postzygotic single-nucleotide mosaicisms in whole-genome sequences of clinically unremarkable individuals
Abstract
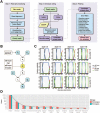
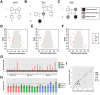
Postzygotic single-nucleotide mutations (pSNMs) have been studied in cancer and a few other overgrowth human disorders at whole-genome scale and found to play critical roles. However, in clinically unremarkable individuals, pSNMs have never been identified at whole-genome scale largely due to technical difficulties and lack of matched control tissue samples, and thus the genome-wide characteristics of pSNMs remain unknown. We developed a new Bayesian-based mosaic genotyper and a series of effective error filters, using which we were able to identify 17 SNM sites from ~80× whole-genome sequencing of peripheral blood DNAs from three clinically unremarkable adults. The pSNMs were thoroughly validated using pyrosequencing, Sanger sequencing of individual cloned fragments, and multiplex ligation-dependent probe amplification. The mutant allele fraction ranged from 5%-31%. We found that C→T and C→A were the predominant types of postzygotic mutations, similar to the somatic mutation profile in tumor tissues. Simulation data showed that the overall mutation rate was an order of magnitude lower than that in cancer. We detected varied allele fractions of the pSNMs among multiple samples obtained from the same individuals, including blood, saliva, hair follicle, buccal mucosa, urine, and semen samples, indicating that pSNMs could affect multiple sources of somatic cells as well as germ cells. Two of the adults have children who were diagnosed with Dravet syndrome. We identified two non-synonymous pSNMs in SCN1A, a causal gene for Dravet syndrome, from these two unrelated adults and found that the mutant alleles were transmitted to their children, highlighting the clinical importance of detecting pSNMs in genetic counseling.
Figures



References
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

