Phospholipid flippase activities and substrate specificities of human type IV P-type ATPases localized to the plasma membrane
- PMID: 25315773
- PMCID: PMC4246107
- DOI: 10.1074/jbc.M114.593012
Phospholipid flippase activities and substrate specificities of human type IV P-type ATPases localized to the plasma membrane
Erratum in
-
Phospholipid flippase activities and substrate specificities of human type IV P-type ATPases localized to the plasma membrane.J Biol Chem. 2016 Oct 7;291(41):21421. doi: 10.1074/jbc.A114.593012. J Biol Chem. 2016. PMID: 27825076 Free PMC article. No abstract available.
Abstract
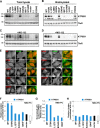
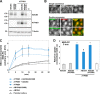
Type IV P-type ATPases (P4-ATPases) are believed to translocate aminophospholipids from the exoplasmic to the cytoplasmic leaflets of cellular membranes. The yeast P4-ATPases, Drs2p and Dnf1p/Dnf2p, flip nitrobenzoxadiazole-labeled phosphatidylserine at the Golgi complex and nitrobenzoxadiazole-labeled phosphatidylcholine (PC) at the plasma membrane, respectively. However, the flippase activities and substrate specificities of mammalian P4-ATPases remain incompletely characterized. In this study, we established an assay for phospholipid flippase activities of plasma membrane-localized P4-ATPases using human cell lines stably expressing ATP8B1, ATP8B2, ATP11A, and ATP11C. We found that ATP11A and ATP11C have flippase activities toward phosphatidylserine and phosphatidylethanolamine but not PC or sphingomyelin. By contrast, ATPase-deficient mutants of ATP11A and ATP11C did not exhibit any flippase activity, indicating that these enzymes catalyze flipping in an ATPase-dependent manner. Furthermore, ATP8B1 and ATP8B2 exhibited preferential flippase activities toward PC. Some ATP8B1 mutants found in patients of progressive familial intrahepatic cholestasis type 1 (PFIC1), a severe liver disease caused by impaired bile flow, failed to translocate PC despite their delivery to the plasma membrane. Moreover, incorporation of PC mediated by ATP8B1 can be reversed by simultaneous expression of ABCB4, a PC floppase mutated in PFIC3 patients. Our findings elucidate the flippase activities and substrate specificities of plasma membrane-localized human P4-ATPases and suggest that phenotypes of some PFIC1 patients result from impairment of the PC flippase activity of ATP8B1.
Keywords: ATPase; Lipid Bilayer; Lipid Transport; Phospholipid; Plasma Membrane.
© 2014 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures



References
-
- Graham T. R. (2004) Flippases and vesicle-mediated protein transport. Trends Cell Biol. 14, 670–677 - PubMed
-
- Pomorski T., Holthuis J. C., Herrmann A., van Meer G. (2004) Tracking down lipid flippases and their biological functions. J. Cell Sci. 117, 805–813 - PubMed
-
- Zwaal R. F., Schroit A. J. (1997) Pathophysiologic implications of membrane phospholipid asymmetry in blood cells. Blood 89, 1121–1132 - PubMed
-
- van den Eijnde S. M., van den Hoff M. J., Reutelingsperger C. P., van Heerde W. L., Henfling M. E., Vermeij-Keers C., Schutte B., Borgers M., Ramaekers F. C. (2001) Transient expression of phosphatidylserine at cell-cell contact areas is required for myotube formation. J. Cell Sci. 114, 3631–3642 - PubMed
-
- Williamson P., Schlegel R. A. (2002) Transbilayer phospholipid movement and the clearance of apoptotic cells. Biochim. Biophys. Acta 1585, 53–63 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

