The long noncoding RNA Neat1 is required for mammary gland development and lactation
- PMID: 25316907
- PMCID: PMC4238351
- DOI: 10.1261/rna.047332.114
The long noncoding RNA Neat1 is required for mammary gland development and lactation
Abstract
The lncRNA Neat1 is an essential architectural component of paraspeckle nuclear bodies. Although cell-based studies identified Neat1-paraspeckles as key regulators of gene expression through retention of hyperdited mRNAs and/or transcription factors, it is unclear under which specific physiological conditions paraspeckles are formed in vivo and whether they have any biological relevance. Herein, we show that paraspeckles are assembled in luminal epithelial cells during mammary gland development. Importantly, genetic ablation of Neat1 results in aberrant mammary gland morphogenesis and lactation defects. We provide evidence that the lactation defect is caused by a decreased ability of Neat1-mutant cells to sustain high rates of proliferation during lobular-alveolar development. This study is the first to assign an important biological function to the lncRNA Neat1 and to link it to the presence of paraspeckles nuclear bodies in vivo.
Keywords: NEAT1; long noncoding RNA; mammary gland development; paraspeckles.
© 2014 Standaert et al.; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures

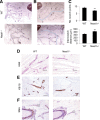
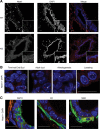
References
-
- Fatica A, Bozzoni I 2014. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet 15: 7–21. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
