The yeast galactose network as a quantitative model for cellular memory
- PMID: 25328105
- PMCID: PMC4252300
- DOI: 10.1039/c4mb00448e
The yeast galactose network as a quantitative model for cellular memory
Abstract
Recent experiments have revealed surprising behavior in the yeast galactose (GAL) pathway, one of the preeminent systems for studying gene regulation. Under certain circumstances, yeast cells display memory of their prior nutrient environments. We distinguish two kinds of cellular memory discovered by quantitative investigations of the GAL network and present a conceptual framework for interpreting new experiments and current ideas on GAL memory. Reinduction memory occurs when cells respond transcriptionally to one environment, shut down the response during several generations in a second environment, then respond faster and with less cell-to-cell variation when returned to the first environment. Persistent memory describes a long-term, arguably stable response in which cells adopt a bimodal or unimodal distribution of induction levels depending on their preceding environment. Deep knowledge of how the yeast GAL pathway responds to different sugar environments has enabled rapid progress in uncovering the mechanisms behind GAL memory, which include cytoplasmic inheritance of inducer proteins and positive feedback loops among regulatory genes. This network of genes, long used to study gene regulation, is now emerging as a model system for cellular memory.
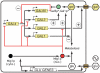
Figures



Similar articles
-
Computational analysis of GAL pathway pinpoints mechanisms underlying natural variation.PLoS Comput Biol. 2021 Sep 27;17(9):e1008691. doi: 10.1371/journal.pcbi.1008691. eCollection 2021 Sep. PLoS Comput Biol. 2021. PMID: 34570755 Free PMC article.
-
Hsp90 mediates the crosstalk between galactose metabolism and cell morphology pathways in yeast.Curr Genet. 2017 Feb;63(1):23-27. doi: 10.1007/s00294-016-0614-2. Epub 2016 May 21. Curr Genet. 2017. PMID: 27209632 Review.
-
Structural bistability of the GAL regulatory network and characterization of its domains of attraction.J Comput Biol. 2012 Feb;19(2):148-62. doi: 10.1089/cmb.2011.0251. J Comput Biol. 2012. PMID: 22300317
-
Pleiotropy and GAL pathway degeneration in yeast.J Evol Biol. 2007 Jul;20(4):1333-8. doi: 10.1111/j.1420-9101.2007.01351.x. J Evol Biol. 2007. PMID: 17584228
-
Regulations of sugar transporters: insights from yeast.Curr Genet. 2013 May;59(1-2):1-31. doi: 10.1007/s00294-013-0388-8. Epub 2013 Mar 1. Curr Genet. 2013. PMID: 23455612 Review.
Cited by
-
Molecular and cellular bases of adaptation to a changing environment in microorganisms.Proc Biol Sci. 2016 Oct 26;283(1841):20161458. doi: 10.1098/rspb.2016.1458. Proc Biol Sci. 2016. PMID: 27798299 Free PMC article. Review.
-
Microbial Adaptation to Enhance Stress Tolerance.Front Microbiol. 2022 Apr 27;13:888746. doi: 10.3389/fmicb.2022.888746. eCollection 2022. Front Microbiol. 2022. PMID: 35572687 Free PMC article. Review.
-
Technological Approach to Mind Everywhere: An Experimentally-Grounded Framework for Understanding Diverse Bodies and Minds.Front Syst Neurosci. 2022 Mar 24;16:768201. doi: 10.3389/fnsys.2022.768201. eCollection 2022. Front Syst Neurosci. 2022. PMID: 35401131 Free PMC article.
-
Probabilistic adaptation in changing microbial environments.PeerJ. 2016 Dec 14;4:e2716. doi: 10.7717/peerj.2716. eCollection 2016. PeerJ. 2016. PMID: 27994963 Free PMC article.
-
Ecological memory of prior nutrient exposure in the human gut microbiome.ISME J. 2022 Nov;16(11):2479-2490. doi: 10.1038/s41396-022-01292-x. Epub 2022 Jul 23. ISME J. 2022. PMID: 35871250 Free PMC article.
References
-
- Davidson EH, Rast JP, Oliveri P, Ransick A, Calestani C, Yuh CH, Minokawa T, Amore G, Hinman V, Arenas-Mena C, Otim O, Brown CT, Livi CB, Lee PY, Revilla R, Rust AG, Pan Z, Schilstra MJ, Clarke PJ, Arnone MI, Rowen L, Cameron RA, McClay DR, Hood L, Bolouri H. Science. 2002;295(5560):1669–1678. - PubMed
-
- Vladimirov N, Sourjik V. Biol Chem. 2009;390(11):1097–1104. - PubMed
-
- Lohr D, Venkov P, Zlatanova J. FASEB J. 1995;9(9):777–787. - PubMed
-
- Zacharioudakis I, Gligoris T, Tzamarias D. Curr Biol. 2007;17(23):2041–2046. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

