At short telomeres Tel1 directs early replication and phosphorylates Rif1
- PMID: 25329891
- PMCID: PMC4199499
- DOI: 10.1371/journal.pgen.1004691
At short telomeres Tel1 directs early replication and phosphorylates Rif1
Abstract
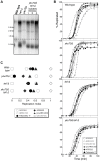
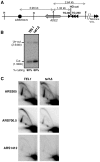
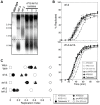
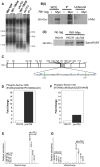
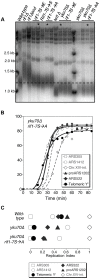
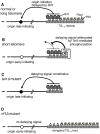
The replication time of Saccharomyces cerevisiae telomeres responds to TG1-3 repeat length, with telomeres of normal length replicating late during S phase and short telomeres replicating early. Here we show that Tel1 kinase, which is recruited to short telomeres, specifies their early replication, because we find a tel1Δ mutant has short telomeres that nonetheless replicate late. Consistent with a role for Tel1 in driving early telomere replication, initiation at a replication origin close to an induced short telomere was reduced in tel1Δ cells, in an S phase blocked by hydroxyurea. The telomeric chromatin component Rif1 mediates late replication of normal telomeres and is a potential substrate of Tel1 phosphorylation, so we tested whether Tel1 directs early replication of short telomeres by inactivating Rif1. A strain lacking both Rif1 and Tel1 behaves like a rif1Δ mutant by replicating its telomeres early, implying that Tel1 can counteract the delaying effect of Rif1 to control telomere replication time. Proteomic analyses reveals that in yku70Δ cells that have short telomeres, Rif1 is phosphorylated at Tel1 consensus sequences (S/TQ sites), with phosphorylation of Serine-1308 being completely dependent on Tel1. Replication timing analysis of a strain mutated at these phosphorylation sites, however, suggested that Tel1-mediated phosphorylation of Rif1 is not the sole mechanism of replication timing control at telomeres. Overall, our results reveal two new functions of Tel1 at shortened telomeres: phosphorylation of Rif1, and specification of early replication by counteracting the Rif1-mediated delay in initiation at nearby replication origins.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
Rif1 phosphorylation site analysis in telomere length regulation and the response to damaged telomeres.DNA Repair (Amst). 2018 May;65:26-33. doi: 10.1016/j.dnarep.2018.03.001. Epub 2018 Mar 7. DNA Repair (Amst). 2018. PMID: 29544213 Free PMC article.
-
Tel1ATM dictates the replication timing of short yeast telomeres.EMBO Rep. 2014 Oct;15(10):1093-101. doi: 10.15252/embr.201439242. Epub 2014 Aug 13. EMBO Rep. 2014. PMID: 25122631 Free PMC article.
-
Rif1 acts through Protein Phosphatase 1 but independent of replication timing to suppress telomere extension in budding yeast.Nucleic Acids Res. 2018 May 4;46(8):3993-4003. doi: 10.1093/nar/gky132. Nucleic Acids Res. 2018. PMID: 29529242 Free PMC article.
-
G-quadruplex binding protein Rif1, a key regulator of replication timing.J Biochem. 2021 Feb 6;169(1):1-14. doi: 10.1093/jb/mvaa128. J Biochem. 2021. PMID: 33169133 Review.
-
Rif1-Dependent Control of Replication Timing.Genes (Basel). 2022 Mar 20;13(3):550. doi: 10.3390/genes13030550. Genes (Basel). 2022. PMID: 35328102 Free PMC article. Review.
Cited by
-
Checkpoint phosphorylation sites on budding yeast Rif1 protect nascent DNA from degradation by Sgs1-Dna2.PLoS Genet. 2023 Nov 13;19(11):e1011044. doi: 10.1371/journal.pgen.1011044. eCollection 2023 Nov. PLoS Genet. 2023. PMID: 37956214 Free PMC article.
-
Rif1 regulates telomere length through conserved HEAT repeats.Nucleic Acids Res. 2021 Apr 19;49(7):3967-3980. doi: 10.1093/nar/gkab206. Nucleic Acids Res. 2021. PMID: 33772576 Free PMC article.
-
Towards the Mechanism of Yeast Telomere Dynamics.Trends Cell Biol. 2019 May;29(5):361-370. doi: 10.1016/j.tcb.2019.01.005. Epub 2019 Feb 11. Trends Cell Biol. 2019. PMID: 30765145 Free PMC article. Review.
-
Rif1 phosphorylation site analysis in telomere length regulation and the response to damaged telomeres.DNA Repair (Amst). 2018 May;65:26-33. doi: 10.1016/j.dnarep.2018.03.001. Epub 2018 Mar 7. DNA Repair (Amst). 2018. PMID: 29544213 Free PMC article.
-
Regulation of DNA Replication through Natural Impediments in the Eukaryotic Genome.Genes (Basel). 2017 Mar 7;8(3):98. doi: 10.3390/genes8030098. Genes (Basel). 2017. PMID: 28272375 Free PMC article. Review.
References
-
- Ferguson BM, Fangman WL (1992) A position effect on the time of replication origin activation in yeast. Cell 68: 333–339. - PubMed
-
- Yamazaki S, Hayano M, Masai H (2013) Replication timing regulation of eukaryotic replicons: Rif1 as a global regulator of replication timing. Trends Genet 29: 449–460. - PubMed
-
- Bianchi A, Shore D (2007) Early replication of short telomeres in budding yeast. Cell 128: 1051–1062. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

