An intramolecular lock facilitates folding and stabilizes the tertiary structure of Streptococcus mutans adhesin P1
- PMID: 25331888
- PMCID: PMC4226092
- DOI: 10.1073/pnas.1413018111
An intramolecular lock facilitates folding and stabilizes the tertiary structure of Streptococcus mutans adhesin P1
Abstract
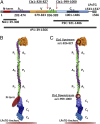
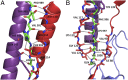
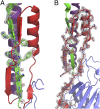
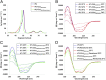
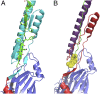
The cariogenic bacterium Streptococcus mutans uses adhesin P1 to adhere to tooth surfaces, extracellular matrix components, and other bacteria. A composite model of P1 based on partial crystal structures revealed an unusual complex architecture in which the protein forms an elongated hybrid alpha/polyproline type II helical stalk by folding back on itself to display a globular head at the apex and a globular C-terminal region at the base. The structure of P1's N terminus and the nature of its critical interaction with the C-terminal region remained unknown, however. We have cocrystallized a stable complex of recombinant N- and C-terminal fragments and here describe a previously unidentified topological fold in which these widely discontinuous domains are intimately associated. The structure reveals that the N terminus forms a stabilizing scaffold by wrapping behind the base of P1's elongated stalk and physically "locking" it into place. The structure is stabilized through a highly favorable ΔG(solvation) on complex formation, along with extensive hydrogen bonding. We confirm the functional relevance of this intramolecular interaction using differential scanning calorimetry and circular dichroism to show that disruption of the proper spacing of residues 989-1001 impedes folding and diminishes stability of the full-length molecule, including the stalk. Our findings clarify previously unexplained functional and antigenic properties of P1.
Keywords: Streptococcus; X-ray crystallography; adhesin; intramolecular lock; protein folding.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Specific binding of a naturally occurring amyloidogenic fragment of Streptococcus mutans adhesin P1 to intact P1 on the cell surface characterized by solid state NMR spectroscopy.J Biomol NMR. 2016 Feb;64(2):153-64. doi: 10.1007/s10858-016-0017-1. Epub 2016 Feb 2. J Biomol NMR. 2016. PMID: 26837620 Free PMC article.
-
An intramolecular interaction involving the N terminus of a streptococcal adhesin affects its conformation and adhesive function.J Biol Chem. 2013 May 10;288(19):13762-74. doi: 10.1074/jbc.M113.459974. Epub 2013 Mar 28. J Biol Chem. 2013. PMID: 23539625 Free PMC article.
-
Characterization of an intermolecular quaternary interaction between discrete segments of the Streptococcus mutans adhesin P1 by NMR spectroscopy.FEBS J. 2020 Jun;287(12):2597-2611. doi: 10.1111/febs.15158. Epub 2019 Dec 17. FEBS J. 2020. PMID: 31782893 Free PMC article.
-
The changing faces of Streptococcus antigen I/II polypeptide family adhesins.Mol Microbiol. 2010 Jul;77(2):276-86. doi: 10.1111/j.1365-2958.2010.07212.x. Epub 2010 May 24. Mol Microbiol. 2010. PMID: 20497507 Free PMC article. Review.
-
Molecular, immunological and functional characterization of the major surface adhesin of Streptococcus mutans.Adv Exp Med Biol. 1992;327:229-41. doi: 10.1007/978-1-4615-3410-5_25. Adv Exp Med Biol. 1992. PMID: 1295342 Review.
Cited by
-
Polymer Adhesin Domains in Gram-Positive Cell Surface Proteins.Front Microbiol. 2020 Nov 26;11:599899. doi: 10.3389/fmicb.2020.599899. eCollection 2020. Front Microbiol. 2020. PMID: 33324381 Free PMC article. Review.
-
Streptococcus Mutans Adhesin Biotypes that Match and Predict Individual Caries Development.EBioMedicine. 2017 Oct;24:205-215. doi: 10.1016/j.ebiom.2017.09.027. Epub 2017 Sep 21. EBioMedicine. 2017. PMID: 28958656 Free PMC article.
-
Specific binding of a naturally occurring amyloidogenic fragment of Streptococcus mutans adhesin P1 to intact P1 on the cell surface characterized by solid state NMR spectroscopy.J Biomol NMR. 2016 Feb;64(2):153-64. doi: 10.1007/s10858-016-0017-1. Epub 2016 Feb 2. J Biomol NMR. 2016. PMID: 26837620 Free PMC article.
-
Backbone NMR resonance assignments for the C terminal domain of the Streptococcus mutans adhesin P1.Biomol NMR Assign. 2023 Dec;17(2):293-299. doi: 10.1007/s12104-023-10158-y. Epub 2023 Oct 21. Biomol NMR Assign. 2023. PMID: 37864759 Free PMC article.
-
Discovery of fibrillar adhesins across bacterial species.BMC Genomics. 2021 Jul 18;22(1):550. doi: 10.1186/s12864-021-07586-2. BMC Genomics. 2021. PMID: 34275445 Free PMC article.
References
-
- Hajishengallis G, Koga T, Russell MW. Affinity and specificity of the interactions between Streptococcus mutans antigen I/II and salivary components. J Dent Res. 1994;73(9):1493–1502. - PubMed
-
- Russell RR. Wall-associated protein antigens of Streptococcus mutans. J Gen Microbiol. 1979;114(1):109–115. - PubMed
-
- Russell MW, Mansson-Rahemtulla B. Interaction between surface protein antigens of Streptococcus mutans and human salivary components. Oral Microbiol Immunol. 1989;4(2):106–111. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

