Isobaric labeling-based relative quantification in shotgun proteomics
- PMID: 25337643
- PMCID: PMC4261935
- DOI: 10.1021/pr500880b
Isobaric labeling-based relative quantification in shotgun proteomics
Abstract
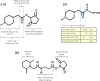
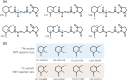
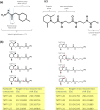
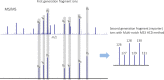
Mass spectrometry plays a key role in relative quantitative comparisons of proteins in order to understand their functional role in biological systems upon perturbation. In this review, we review studies that examine different aspects of isobaric labeling-based relative quantification for shotgun proteomic analysis. In particular, we focus on different types of isobaric reagents and their reaction chemistry (e.g., amine-, carbonyl-, and sulfhydryl-reactive). Various factors, such as ratio compression, reporter ion dynamic range, and others, cause an underestimation of changes in relative abundance of proteins across samples, undermining the ability of the isobaric labeling approach to be truly quantitative. These factors that affect quantification and the suggested combinations of experimental design and optimal data acquisition methods to increase the precision and accuracy of the measurements will be discussed. Finally, the extended application of isobaric labeling-based approach in hyperplexing strategy, targeted quantification, and phosphopeptide analysis are also examined.
Keywords: TMT; iTRAQ; isobaric labeling; isobaric tags; isobaric tags for relative and absolute quantification; mass spectrometry; quantitative proteomics; tandem mass tags.
Figures



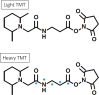
References
-
- Wu C. C.; MacCoss M. J.; Howell K. E.; Matthews D. E.; Yates J. R. III Metabolic labeling of mammalian organisms with stable isotopes for quantitative proteomic analysis. Anal. Chem. 2004, 76, 4951–4959. - PubMed
-
- Ong S. E.; Blagoev B.; Kratchmarova I.; Kristensen D. B.; Steen H.; Pandey A.; Mann M. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 2002, 1, 376–386. - PubMed
-
- Gygi S. P.; Rist B.; Gerber S. A.; Turecek F.; Gelb M. H.; Aebersold R. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 1999, 17, 994–999. - PubMed
-
- Hsu J. L.; Huang S. Y.; Chow N. H.; Chen S. H. Stable-isotope dimethyl labeling for quantitative proteomics. Anal. Chem. 2003, 75, 6843–6852. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

