Quantifying distortions in two-photon remote focussing microscope images using a volumetric calibration specimen
- PMID: 25339910
- PMCID: PMC4189333
- DOI: 10.3389/fphys.2014.00384
Quantifying distortions in two-photon remote focussing microscope images using a volumetric calibration specimen
Abstract
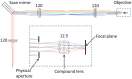
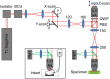
Remote focussing microscopy allows sharp, in-focus images to be acquired at high speed from outside of the focal plane of an objective lens without any agitation of the specimen. However, without careful optical alignment, the advantages of remote focussing microscopy could be compromised by the introduction of depth-dependent scaling artifacts. To achieve an ideal alignment in a point-scanning remote focussing microscope, the lateral (XY) scan mirror pair must be imaged onto the back focal plane of both the reference and imaging objectives, in a telecentric arrangement. However, for many commercial objective lenses, it can be difficult to accurately locate the position of the back focal plane. This paper investigates the impact of this limitation on the fidelity of three-dimensional data sets of living cardiac tissue, specifically the introduction of distortions. These distortions limit the accuracy of sarcomere measurements taken directly from raw volumetric data. The origin of the distortion is first identified through simulation of a remote focussing microscope. Using a novel three-dimensional calibration specimen it was then possible to quantify experimentally the size of the distortion as a function of objective misalignment. Finally, by first approximating and then compensating the distortion in imaging data from whole heart rodent studies, the variance of sarcomere length (SL) measurements was reduced by almost 50%.
Keywords: cardiac imaging; distortion; remote focussing microscopy.
Figures










References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

